29 March 2022: Database Analysis
Bioinformatics Identification of Candidate Biomarkers in Endomyocardial Biopsy and Peripheral Blood for Cardiac Allograft Rejection
Kang Luo1ABCDEFG, Lin Li2BCDE, Mingyao Meng2DEFG, Yan Chen1BC, Zongliu Hou2EFDOI: 10.12659/AOT.935488
Ann Transplant 2022; 27:e935488
Abstract
BACKGROUND: Cardiac allograft rejection is still a crucial barrier to achieving satisfactory outcomes after surgery. In this study, we propose to find candidate biomarkers from endomyocardial biopsy (EMB) and peripheral blood (PB) samples for efficient diagnosis and treatment of cardiac allograft rejection.
MATERIAL AND METHODS: Microarray datasets were obtained from the Gene Expression Omnibus (GEO). Differentially expressed genes (DEGs) of cardiac allograft rejection patients and control subjects from EMB and PB samples were screened using the online tool GEO2R. Gene ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway enrichment analysis of all samples’ DEGs were performed with the DAVID online tool. Protein–protein interaction (PPI) networks were constructed and visualized using Cytoscape and the top 10 hub genes were selected. Finally, the most highly enriched GO and KEGG pathways of the top 10 hub genes were determined.
RESULTS: A total of 57 502 genes from EMB samples and 131 624 genes from PB samples were identified. Gene characteristics and enrichment analysis indicated that both EMB and PB samples contained DEGs involved in antigen presentation, immune cells activation, inflammatory process, and cellular injuries. In EMB samples, there were some DEGs related to heart tissue injury and cardiac malfunction. Moreover, DEGs that regulates hypoxia-induced factors and erythrocyte function in response of ischemia and hypoxia stress were present in PB samples but were absent in EMB samples.
CONCLUSIONS: The screened differentially expressed genes (DEGs) from EMB and PB samples of patients with cardiac graft rejection are potential candidate biomarkers of diagnosis and treatment.
Keywords: Graft Rejection, Heart Transplantation, Allografts, Biopsy, Computational Biology, Humans, Tissue Donors
Background
Heart transplantation is one of the most effective options to treat patients with end-stage heart disease. However, acute immune rejection is still an intractable obstacle for prolonging survival and promoting quality of life after surgery [1]. To solve this urgent issue, the first and most crucial step is to find biomarkers that can be used in rapid and accurate diagnosis for early-stage immune rejection disorder. Additionally, finding efficient therapeutic targets for alleviating rejection injury before the occurrence irreversible graft failure and multiple organ failure is also important.
In clinical practice, endomyocardial biopsy (EMB) is the criterion standard to identify acute cardiac rejection [2]. However, drawbacks of EMB have been realized since this technology has been initially applied decades ago. On the one hand, EMB is an invasive procedure that inevitably causes complications, including cardiac tamponade, arrhythmias, severe heart internal-structure injury, bleeding, and thrombosis [3]. On the other hand, EMB is mainly based on the histopathologic interpretation, which is relatively insensitive to detect early rejection [4]. To find a better way to diagnose and intervene in heart transplant rejection, in recent years, researchers have been searching for find biomarkers that can improve the accuracy of EMB and make it more suitable for early detection of rejection [5,6]. For example, it was found that a set of microRNAs are expressed differently in EMB samples from patients with heart transplant rejection in earlier stage [7]. Meanwhile, other researchers are committed to the development of minimally invasive and non-invasive cardiac allograft rejection monitoring, including traditional imaging diagnosis, intramyocardial electrocardiogram, molecular imaging, and testing biomarkers in peripheral blood (PB). Among the minimally-invasive and non-invasive methods currently used, testing of PB samples has been regarded as the most repeatable, convenient, and economical method for diagnosis of rejection [8]. It can determine whether the type of rejection is acute cellular rejection (ACR), antibody-mediated rejection (AMR), or mixed. And novel biomarkers such as cell-free DNA in peripheral blood enable this test more sensible for diagnosis [4]. However, limitations still exist, as the peripheral blood test has poor ability to distinguish rejection and infection from immunological stability [8,9], so further investigations to identify biomarkers of cardiac allograft rejection from EMB and PB are warranted.
In this study, we analyzed microarray data of rejection and non-rejection heart transplantation recipients from EMB and PB samples by bioinformatics methods, which were downloaded from the Gene Expression Omnibus (GEO) database. Then, we screened out the differentially expressed genes (DEGs) from the 2 types of samples, displayed their significant Gene Ontology (GO) functional annotation analysis and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway enrichment, and constructed the protein–protein interaction (PPI) networks. Through further analysis, the hub genes of EMB and PB samples and their specific bio-function in cardiac graft rejection was discovered. By performing in-depth analysis and comparing characteristics of the DEGs and pivotal roles they likely play in cardiac graft rejection, we selected the candidate biomarkers of the 2 sample types that possess huge potential value to improve future cardiac allograft rejection diagnosis and treatment.
Material and Methods
MICROARRAY SOURCE:
By searching public resources in the Gene Expression Omnibus (GEO) database (
DATA PROCESSING AND DEG IDENTIFICATION:
The raw microarray data including EMB and PB samples were obtained from the GEO database. After filtering out chronic rejection cases, data of both samples were analyzed separately by the online tool GEO2R (http://www.ncbi.nlm.nih.gov/geo/geo2r) to screen out different genes expressed between rejection and non-rejection samples. We downloaded data files analyzed by GEO2R and we used the OmicStudio tools (https://www.omicstudio.cn/tool) to show the volcano plots of genes expressed in EMB and PB samples from rejection and non-rejection groups (Figure 1). Among these genes, those that meet the standard of P value <0.05 and |logFC| ≥1.0 were regarded as significant differentially expressed genes (DEGs). According to the absolute value of logFC ranked from high to low, the top 10 DEGs from both samples were selected (Tables 1, 2).
ENRICHMENT ANALYSES OF DEGS:
DEGs expression profiles were submitted to the DAVID online tool (Database for Annotation, Visualization and Integrated Discovery, version 6.8, http://david.ncfcrf.gov/) for analysis. To assess the significance of the biological function of DEGs, GO and KEGG pathway annotation were performed for EMB and PB samples. P<0.05 and gene counts ≥10 were set as the threshold for EMB samples. In the case of PB samples, P<0.05 and gene counts ≥5 were considered statistically significant (Tables 3, 4).
PPI NETWORK CONSTRUCTION AND HUB GENES IDENTIFICATION:
Protein–protein interaction (PPI) networks of DEGs for EMB and PB samples were constructed and visualized with Cytoscape software (version 3.8.2). A confidence score cutoff >0.4 was set as the standard of significance for PPI pairs (Figure 2). The top 10 hub genes of each PPI network were selected and entered into CytoHubba, a plugin in Cytoscape software, according to the degree of nodes rank (≥10) from high to low (Figure 3). Then, we analyzed GO and KEGG pathway enrichment of the top 10 hub genes (Tables 5, 6).
Results
IDENTIFICATION OF THE DEGS:
Genes expressed in EMB and PB samples of patients with and without rejection after heart transplantation were obtained by submitting microarray data to the GEO2R analyzer, resulting in 57 502 genes from EMB samples and 131 624 genes from PB samples. The results were demonstrated via volcano plot (Figure 1) where red squares represented upregulated DEGs (53 genes for EMB and 164 for PB) (logFC ≥1, P<0.05), blue triangles represent downregulated DEGs (88 genes for EMB and 118 for PB) (logFC ≤−1, P<0.05), and the gray circles represent non-differential expression genes (57 361 genes for EMB and 131 342 for PB) (−1 <logFC <1, P≥0.05). Among all of DEGs, the top 10 for EMB and PB samples are shown in Tables 1 and 2, respectively
BIOFUNCTIONAL ENRICHMENT ANALYSIS OF DEGS:
The GO and KEGG pathway enrichment analyses of DEGs were performed using the DAVID online tool. In EMB samples, there were 19 GO and 4 KEGG pathway categories mostly enriched. For GO, cellular component (CC) was the most significant category, including protein binding, protein homodimerization activity, identical protein binding, receptor binding, integral component of membrane, Golgi apparatus, plasma membrane, extracellular exosome, integral component of plasma membrane, membrane, extracellular region, extracellular space, immune response, and Golgi membrane. Biological process (BP) is the second category, consisting of innate immune response, cell surface, signal transduction, T cell receptor signaling pathway, and inflammatory response. For molecular function (MF), protein binding, protein homodimerization activity, identical protein binding, and receptor binding were enriched. Four categories of KEGG pathway in EMB were Antigen processing and presentation, Herpes simplex infection, Cell adhesion molecules (CAMs), and Influenza A (Table 3). However, in PB samples, DEGs were only significantly enriched in GO, which contained 4 categories of CC (cytosol, membrane, mitochondrial outer membrane, and cytoplasm), 4 categories of BP (protein targeting to Golgi, Nuclear-transcribed mRNA catabolic process, Odontogenesis of dentin-containing tooth, and phosphatidylinositol biosynthetic process), and 3 categories of MF (protein binding, ubiquitin conjugating enzyme binding, and protein homodimerization activity) (Table 4).
CONSTRUCTION OF THE PPI NETWORKS AND HUB GENES ANALYSIS:
DEGs for EMB and PB samples were imported independently into Cytoscape software (version 3.8.2) for protein–protein interaction analysis. After setting a confidence score cutoff >0.4 as the threshold value and removing proteins without obvious connection with majority members, PPI networks for each group were constructed and visualized using Cytoscape software (Figure 2). Next, the top 10 hub genes from PPI networks of EMB and PB samples were identified with degrees ≥10, which contained HLA-DQA1, CD74, TYROBP, HLA-B, HLA-C, CCL5, HLA-DRA, B2M, and PSMB9 in EMB samples and SLC4A1, NEDD4L, PIKFYVE, AHSP, SIAH2, ALAS2, RNF14, SNCA, UBA6, and GYPB in PB samples (Figure 3). The top 10 hub genes from EMB were obviously enriched in GO, including protein binding, integral component of membrane, plasma membrane, membrane, immune response, cell surface, extracellular exosome, and T cell receptor signaling pathway. In the KEGG pathway, antigen processing and presentation, herpes simplex infection, and cell adhesion molecules (CAMs) were included. In PB samples, the most enriched GO by top 10 hub genes was protein binding (MF) then followed by cytoplasm, cytosol, intracellular (CC), protein polyubiquitination B, intracellular signal transduction, phosphatidylinositol biosynthetic process, and negative regulation of apoptotic process (BP) (Tables 5, 6).
Discussion
Diagnosing cardiac allograft rejection is mostly based on EMB and PB testing. EMB possesses relatively higher accuracy, but it requires special equipment and skilled physicians, and it is associated with some complications in patients [10]. On the contrary, PB is the most convenient minimally invasive test for cardiac allograft rejection diagnosis and can be performed repeatedly, but its results can be easily influenced by infection or other pathological conditions [11]. Therefore, comprehensively considering the research and analysis of gene expression changes in EMB and PB can potentially improve the diagnosis and treatment of allogeneic cardiac rejection.
In this study, the first step completed was that the top 10 DEGs from EMB and PB samples were selected as representatives, and functions of these genes were annotated. In EMB samples, most of the top 10 DEGs were related to antigen presentation, immune cell activation, and inflammatory process. Genes closely related to immune functions were: IGHG3 (Immunoglobulin Heavy Constant Gamma 3), IGL@ (Immunoglobulin Lambda Locus), CD2, SCYA5 (C-C Motif Chemokine Ligand 5, CCL5), CCL18 (C-C Motif Chemokine Ligand 18), UBD (Ubiquitin D), CSPG2 (Versican), COPG (COPI Coat Complex Subunit Gamma), and NPPB (Natriuretic Peptide B). For example, upregulation of IGHG3 and IGL@ reflects B lymphocytes (B cells) activation, proliferation, differentiation, and immunoglobulins secretion in humoral immunity [12,13]. CD2 is a costimulatory gene highly expressed on activated T and NK cell surfaces [14]. SCYA5 can be released by activated CD8+ T cell, vessel smooth muscle cells, and endothelial cells to induce immune and inflammation reaction by binding to receptors like CCR1, CCR3, and CCR5 [15,16]. CCL18 (C-C Motif Chemokine Ligand 18) not only activates T cells in cellular immune rejection but also promotes immune function of B cells and macrophages in humoral immune response [17]. Additionally, UBD (Ubiquitin D) is a T cell pro-inflammatory cytokine that promotes T cells to excrete TNF-α, inducing cellular apoptosis [18]. Moreover, CSPG2 and COPG are cellular component and function executor proteins, both of which are expressed in endothelial cells, and their increased expression can indicate NK cell response and endothelial activation [19,20]. There are also some genes reflecting heart injury and functional compromise. NPPB mediates cardio-renal homeostasis by regulating the extracellular fluid volume and maintaining the fluid-electrolyte balance, and it increases in patients with congestive heart failure (CHF). SCYA5, mentioned above, is involved in myocardial infarction [21]. In summary, these critical gene changes may reflect the immune landscape and tissue injuries in the EBM samples, and finally reveal tissue pathological conditions, if present.
Next, gene changes in PB groups were evaluated. In PB groups, apart from immune recognition, inflammatory process, and cellular injury, the top 10 DEGs also reflect systematic hypoxia and specific corresponding change of erythrocytes, and these findings are different from those in EBM samples. Firstly, the main genes involved in immune reaction, inflammatory, and organ injury are PBX1 (Pre-B-Cell Leukemia Homeobox 1), IFI27 (interferon Alpha Inducible Protein 27), and RNF182 (ring finger protein 182). Specifically, PBX1 upregulation may indicate B cell activation and expansion [22], and the increased expression of IFI27 and RNF182 not only reflects innate immune response, but also indicates injuries to vessels and myocardium induced by inflammation and ischemia-reperfusion [23,24]. Most importantly, genes indicating hypoxia situation and corresponding changes of erythrocytes were found in PB samples. The expressions of FECH (Ferrochelatase) and BPGM (Bisphosphoglycerate Mutase) usually increase in hypoxia, which then activate and up-regulate hypoxia-responsive genes like hypoxia-induced factor-1 (HIF-1) to compensate for oxygen deficiency [25]. BPGM can decrease the oxygen affinity of hemoglobin in red blood cells, thus helping to compensate for tissue oxygen insufficiency [26]. Meanwhile, XK (X-Linked Kx Blood Group) and HEMGN (Hemogen) are positively associated with erythrocyte and hematopoietic cell generation, proliferation, and differentiation [27,28], which can be stimulated by hypoxia and inflammation [29–31]. These changes in hypoxia- and inflammation-related genes may indicate hypoxia and inflammation in the transplanted heart and can be used as biomarkers.
Next, through enrichment analyses of DEGs from EMB and PB samples, we selected the most centralized GO and KEGG pathway. The top 10 hub genes of PPI networks from EMB and PB were screened out and their involvement in GO and KEGG pathways were analyzed. Firstly, in the EMB group, the top 10 hub genes were enriched in GO of integral component of membrane, plasma membrane, and cell surface as CC (cellular component), immune response, T cell receptor signaling pathway, innate immune response, and inflammatory response as BP (biological process), as well as KEGG pathway of antigen processing and presentation and cell adhesion molecules (CAMs). These categories of GO and KEGG pathways were also reported in recent organ transplantation rejection research [32]. Hub genes, which were enriched in these categories of GO and KEGG pathways mentioned above, also play roles in antigen presentation, immune cell activation, and inflammation. Specifically, HLA class I heavy chain paralogues or component (HLA-B, HLA-C and B2M (Beta-2-Microglobulin) and HLA class II alpha chain paralogues and component (HLA-DQA1, HLA-DRA and CD74) play important roles in the stage of immune recognition, which can recognize exogenous antigen and present them to immune cells like T cells and NK cells [33–36]. TYROBP (Transmembrane Immune Signaling Adaptor) and PSMB9 (Proteasome 20S Subunit Beta 9) participates in the immune activation and immune response processes. They are receptors located on some immune cells, including NK cells, T cells, B cells, dendritic cells, and macrophages, and they are responsible for the activation and regulation of these immune cells [37,38]. In the PB group, categories of mostly enriched GO terms in top 10 hub genes included protein binding (MF), intracellular signal transduction, and negative regulation of apoptotic process (BP). The hub genes enriched in these GO terms above are mainly involved in systematic inflammation, hypoxic stress, cellular injury, and erythrocyte changes. In inflammation, RNF14 (Ring Finger Protein 14) serves as a T cell factor/lymphoid enhancer factor protein in the cellular immune response. Meanwhile, PIKFYVE (Phosphoinositide Kinase and FYVE-Type Zinc Finger Containing) loss in macrophages can damage the lysosomal degradative function, which promotes systemic inflammation [39]. SIAH2 (Siah E3 Ubiquitin Protein Ligase 2) and UBA6 (Ubiquitin Like Modifier Activating Enzyme 6) are involved in hypoxic stress and cellular injury. SIAH2 upregulation can promote stability of HIF-1 responding to hypoxia stress against cell apoptosis [40], while UBA6 downregulation can exacerbate cellular apoptosis in organ ischemia [41]. Finally, ALAS2 (5′-Aminolevulinate Synthase 2), SLC4A1 (Solute Carrier Family 4 Member 1 (Diego Blood Group)), GYPB (GlycophornsB), and AHSP (Alpha Hemoglobin Stabilizing Protein) are genes regulated to bio-activities. Upregulation of these genes can compensate for organ function impairment and may be a marker of the early stages of rejection. In detail, ALAS2 is the specific enzyme located on erythroid mitochondria, SLC4A1 is the integral membrane glycoprotein of the erythrocyte membrane, and GYPB (Glycophorins B) is major sialoglycoproteins of human erythrocytes. Upregulation of these 3 genes can indicate erythrocyte proliferation and activation [42–44]. AHSP (alpha hemoglobin stabilizing protein) is a molecular chaperone which can bind nascent apo-α-globin in the erythroid cell for its development [45], and can also promote NO function of vasodilatation to regulate blood pressure by binding α-globin in arteriolar endothelial cells [46].
In this study, we applied bioinformatics analysis to identify cardiac allograft rejection DEGs and relevant GO, KEGG Pathway, PPI networks, and hub genes from EMB and BP samples. The most significant genes and enrichment were analyzed in-depth as representations. We found that some DEGs from EMB and PB samples are involved in antigen presentation, immune cells activation, inflammatory process, and cellular injuries such as HLA-B, HLA-C, CD74, IGL@, SCYA5, CCL18, IFI27, RNF14, UBD, and UBA6. These genes can be used as biomarkers to identify specific kinds of or the degree of immune activation, so they can guide us to take some action to palliate immune injury before deterioration. In EMB samples, we screened out some genes related to heart tissue injury and cardiac function compromise, including SCYA5 and NPPB. The changed expression of these genes may indicate the deterioration of a cardiac graft, which can remind physicians not only to deal with immune rejection problems but also to address cardiac malfunction [47,48]. In the PB samples, genes containing FECH, BPGM, SIAH2, HEMGN, GYPB ALAS2, SLC4A1, and AHSP regulate hypoxia-induced factor-1 and erythrocyte’s protein synthesis, proliferation, and oxygen affinity as a compensatory mechanism. Changes in expression of these genes sometimes precede the apparent symptoms of immune rejection and cardiac dysfunction. In this way, they can be used as diagnostic early-stage biomarkers and therapeutic targets. Our study suggests how to interpret the nuances of biomarkers of cardiac rejection between EMB and PB samples and to consider the results together. In addition, unlike previous cardiac allograft rejection bioinformatic studies in which only single-center specimens were used or a single kind of sample was analyzed [7,32,49], our present results are more comprehensive. However, a limitation of the present study is that we did not perform in vitro or in vivo experiments on DEGs and their function relating to cardiac graft rejection. Hence, further mechanism studies of DEGs should be conducted to identify biomarkers in a more specific and accurate way to provide an experimental basis for the diagnosis and treatment of cardiac graft rejection at the very early stage.
Conclusions
In this bioinformatics analysis, DEGs from EMB and PB samples of patients with and without cardiac graft rejection were screened and have potential to be candidate biomarkers of rejection diagnosis and treatment. These biomarkers, including CD2, SCYA5, CCL18, UBD, NPPB, HLA-B, HLA-C, PSMB9, B2M, and TYROBP in EMB samples, and PBX1, IFI27, FECH, BPGM, HEMGN, RNF14, PIKFYVE, SIAH2, UBA6, ALAS2, SLC4A1, GYPB, and AHSP in PB samples, can help physicians to better manage cardiac rejection problems. However, more samples should be included and further fundamental investigations should be conducted to identify the translational value of these biomarkers.
Figures
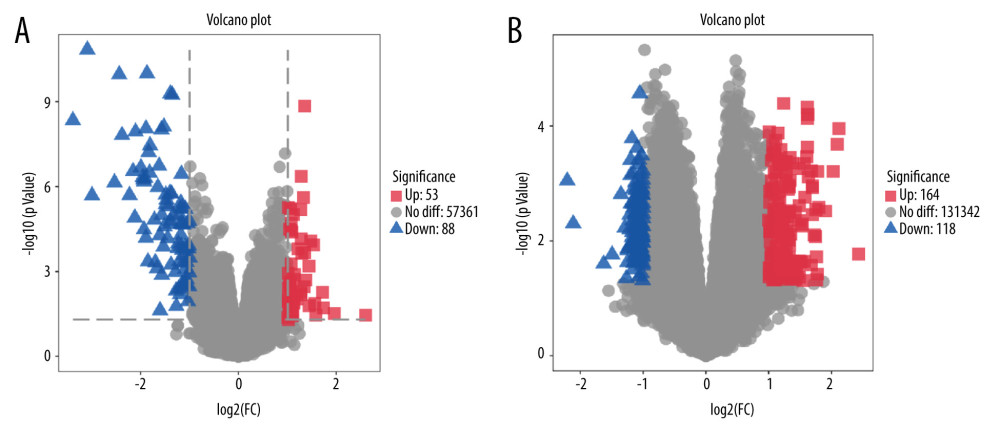 Figure 1. Differential expression of genes from endomyocardial biopsy and peripheral blood samples(A) Volcano plot of 57 502 genes expressed differentially in endomyocardial biopsy of patients with and without cardiac rejection. (B) Volcano plot of 131 624 genes expressed differentially in peripheral blood of patients with and without cardiac rejection. Red plots represent significantly upregulated genes (logFC ≥1, P value <0.05), blue plots are significantly downregulated genes (logFC ≤−1, P value <0.05), while the gray ones demonstrate genes with non-differential expression. Volcano plot graphs were created using the OmicStudio tools available at https://www.omicstudio.cn/tool.
Figure 1. Differential expression of genes from endomyocardial biopsy and peripheral blood samples(A) Volcano plot of 57 502 genes expressed differentially in endomyocardial biopsy of patients with and without cardiac rejection. (B) Volcano plot of 131 624 genes expressed differentially in peripheral blood of patients with and without cardiac rejection. Red plots represent significantly upregulated genes (logFC ≥1, P value <0.05), blue plots are significantly downregulated genes (logFC ≤−1, P value <0.05), while the gray ones demonstrate genes with non-differential expression. Volcano plot graphs were created using the OmicStudio tools available at https://www.omicstudio.cn/tool. 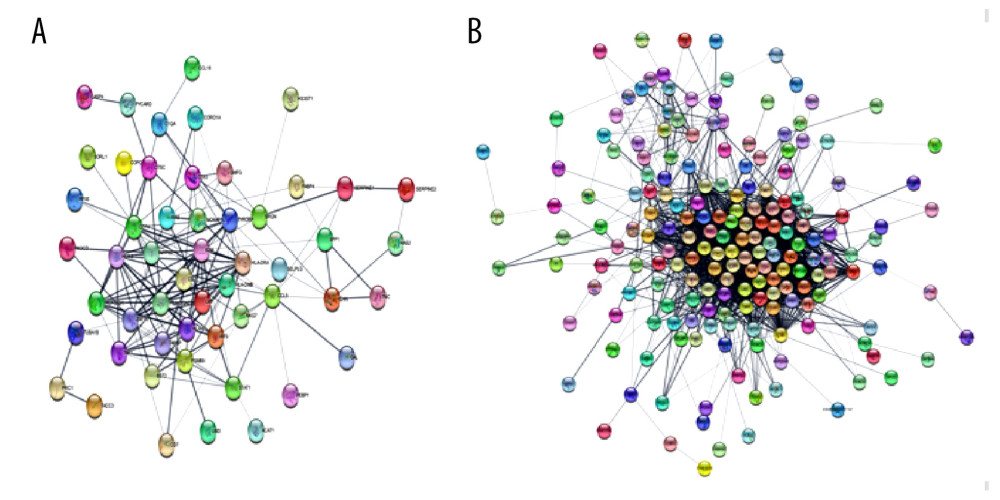 Figure 2. PPI Networks of significant DEGs in endomyocardial biopsy and peripheral blood sample constructed with Cytoscape 3.8.2 software(A) Significant DEGs PPI networks of myocardium. (B) Significant DEGs PPI networks of peripheral blood. Nodes represent genes. Edges represent the interaction among proteins. the structure of proteins can be checked in nodes. Colors of edges represent the interaction between the proteins proved by different kinds of evidence. PPI – protein–protein interaction.
Figure 2. PPI Networks of significant DEGs in endomyocardial biopsy and peripheral blood sample constructed with Cytoscape 3.8.2 software(A) Significant DEGs PPI networks of myocardium. (B) Significant DEGs PPI networks of peripheral blood. Nodes represent genes. Edges represent the interaction among proteins. the structure of proteins can be checked in nodes. Colors of edges represent the interaction between the proteins proved by different kinds of evidence. PPI – protein–protein interaction. 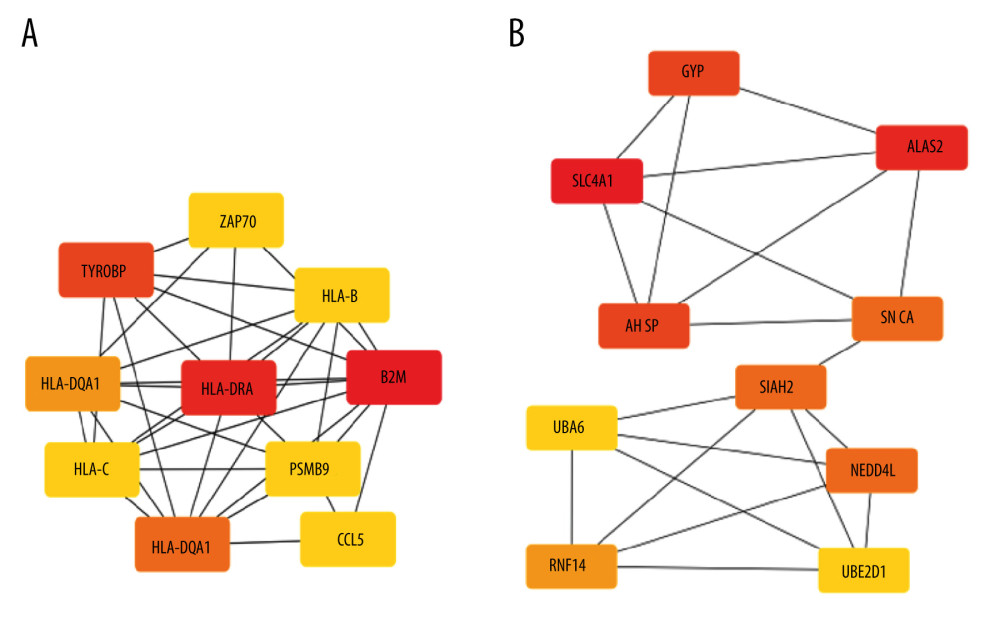 Figure 3. Networks of top 10 hub genes with connective degree of ≥10 achieved with CytoHubba, a plugin in Cytoscape(A) Ten hub genes of DEGs from endomyocardial biopsy samples. (B) Ten hub genes of DEGs from peripheral blood. The depth of color of each gene reflects its degree of connection.
Figure 3. Networks of top 10 hub genes with connective degree of ≥10 achieved with CytoHubba, a plugin in Cytoscape(A) Ten hub genes of DEGs from endomyocardial biopsy samples. (B) Ten hub genes of DEGs from peripheral blood. The depth of color of each gene reflects its degree of connection. Tables
Table 1. Top 10 differentially expressed genes (DEGs) from endomyocardial biopsy samples.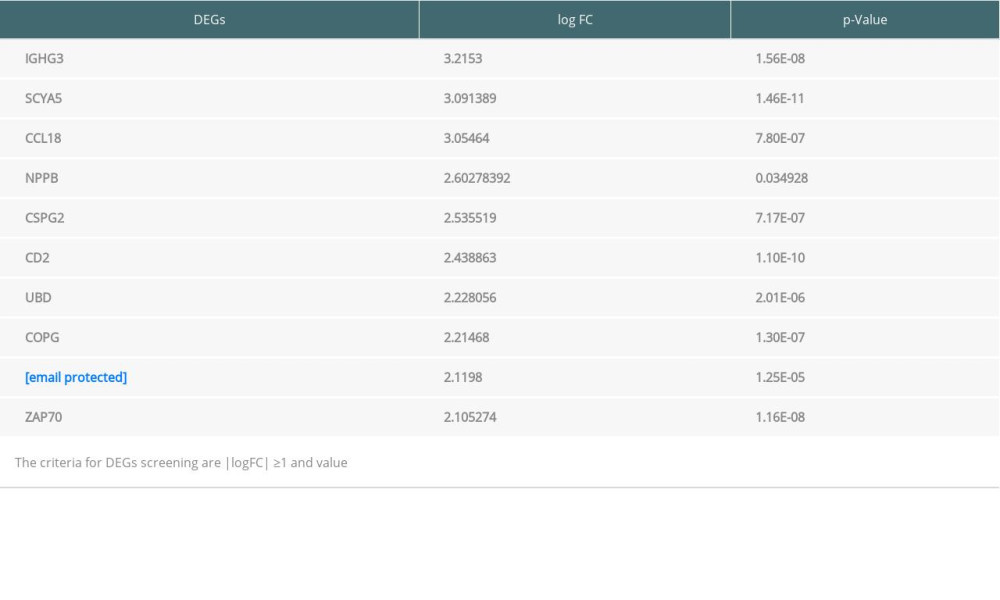 Table 2. Top 10 differentially expressed genes (DEGs) from peripheral blood samples.
Table 2. Top 10 differentially expressed genes (DEGs) from peripheral blood samples.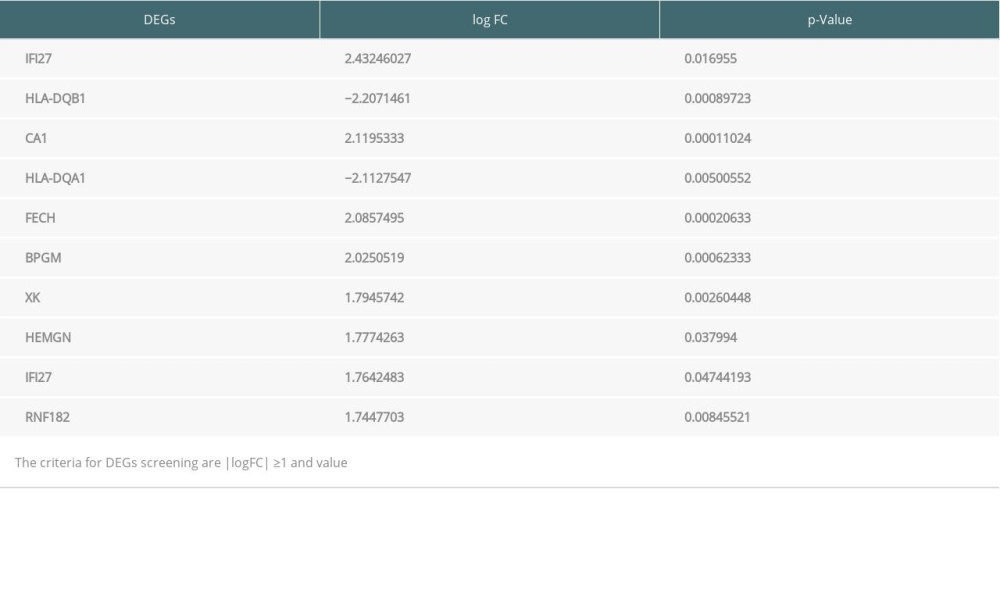 Table 3. Enrichment of Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) analysis of significance for differentially expressed genes (DEGs) from endomyocardial biopsy samples.
Table 3. Enrichment of Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) analysis of significance for differentially expressed genes (DEGs) from endomyocardial biopsy samples.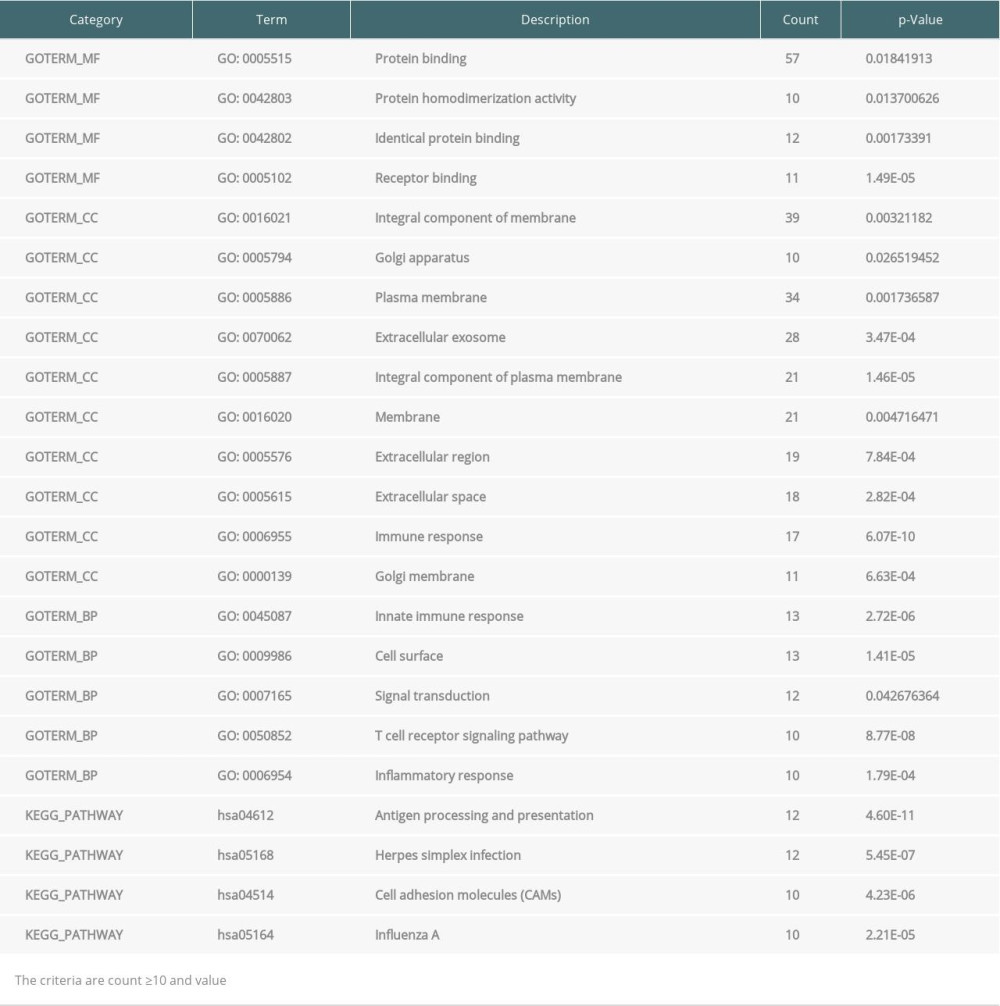 Table 4. Enrichment of Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) analysis of significance for differentially expressed genes (DEGs) from peripheral blood samples.
Table 4. Enrichment of Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) analysis of significance for differentially expressed genes (DEGs) from peripheral blood samples.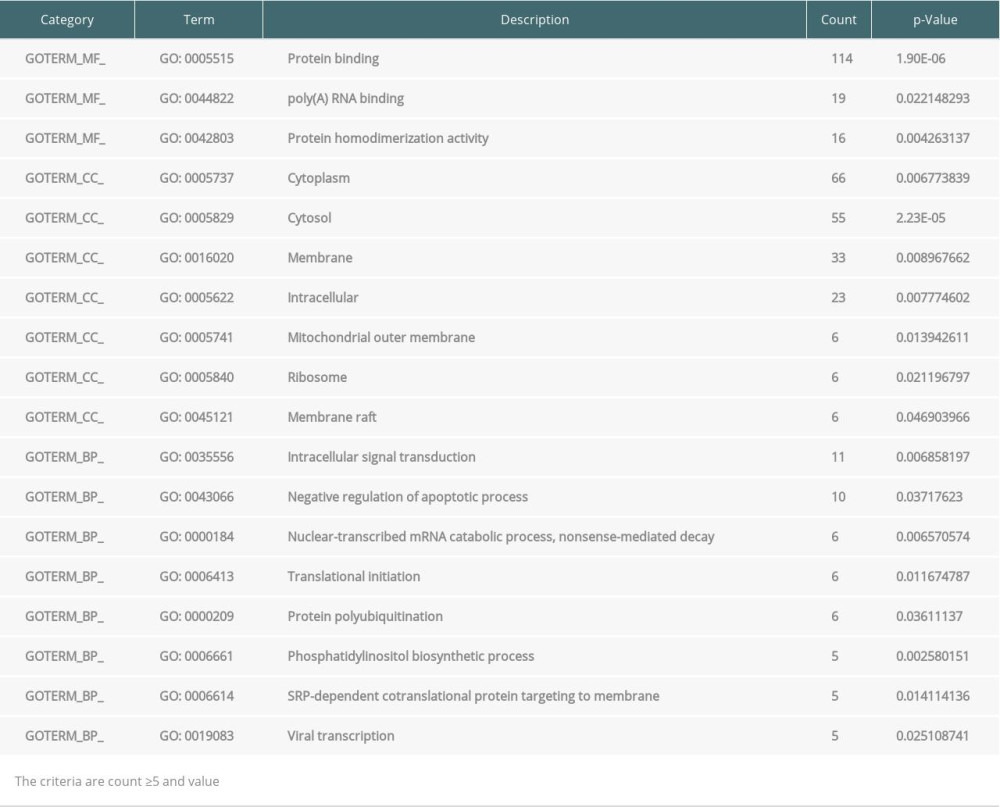 Table 5. Top 10 hub genes from endomyocardial biopsy samples enriched in Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG).
Table 5. Top 10 hub genes from endomyocardial biopsy samples enriched in Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG).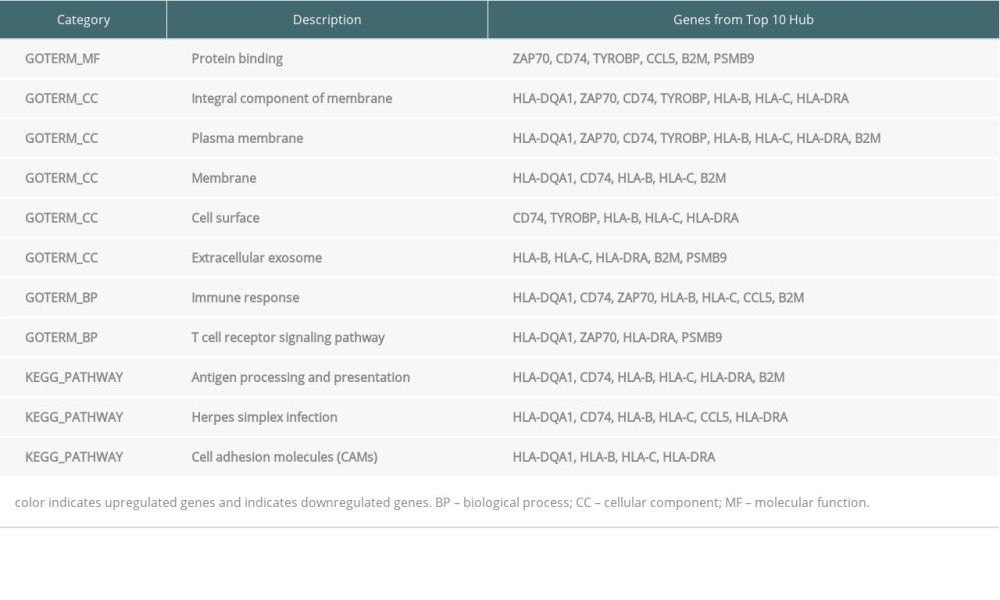 Table 6. Top 10 hub genes from peripheral blood samples enriched in Gene Ontology (GO).
Table 6. Top 10 hub genes from peripheral blood samples enriched in Gene Ontology (GO).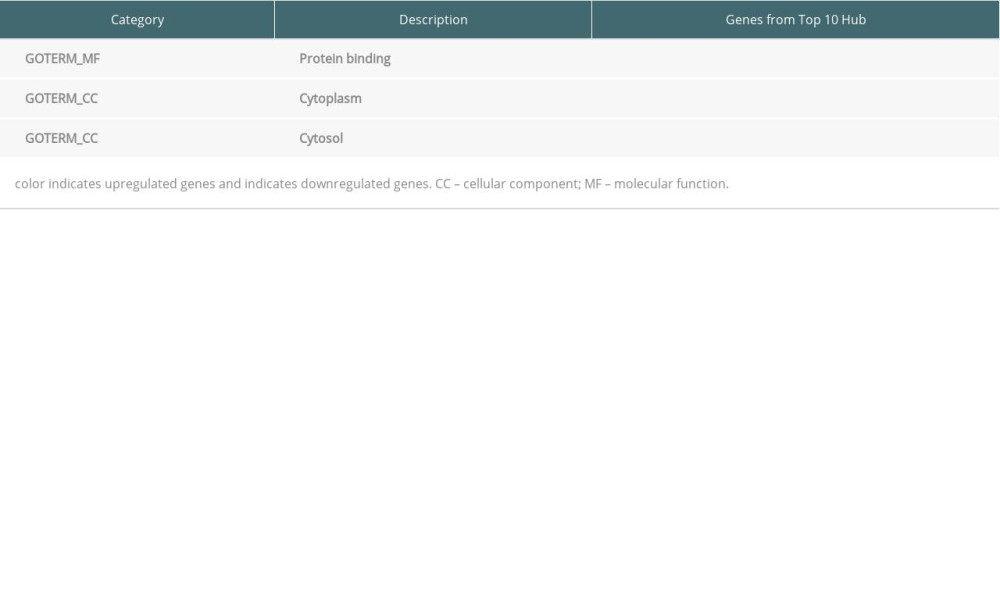
References
1. Weber BN, Kobashigawa JA, Givertz MM, Evolving areas in heart transplantation: JACC Heart Fail, 2017; 5(12); 869-78
2. Patel JK, Kittleson M, Kobashigawa JA, Cardiac allograft rejection: Surgeon, 2011; 9(3); 160-67
3. Shen N, Yan Z, He PA study of the hereditary susceptibility of HLA-DQA1 to essential hypertension, athrothrombotic brain infarction and lacunar stroke: Zhonghua Yi Xue Za Zhi, 2001; 81(6); 352-55 [in Chinese]
4. Agbor-Enoh S, Shah P, Tunc I, Cell-free DNA to detect heart allograft acute rejection: Circulation, 2021; 143(12); 1184-97
5. Halloran PF, Potena L, Van Huyen JD, Building a tissue-based molecular diagnostic system in heart transplant rejection: The heart Molecular Microscope Diagnostic (MMDx) System: J Heart Lung Transplant, 2017; 36(11); 1192-200
6. Parkes MD, Aliabadi AZ, Cadeiras M, An integrated molecular diagnostic report for heart transplant biopsies using an ensemble of diagnostic algorithms: J Heart Lung Transplant, 2019; 38(6); 636-46
7. Novakova T, Machackova T, Novak J, Identification of a diagnostic set of endomyocardial biopsy microRNAs for acute cellular rejection diagnostics in patients after heart transplantation using next-generation sequencing: Cells, 2019; 8(11); 1400
8. Dandel M, Hetzer R, Non-invasive cardiac allograft rejection surveillance: Reliability and clinical value for prevention of heart failure: Heart Fail Rev, 2021; 26(2); 319-36
9. Wong MS, Boucek R, Kemna M, Immune cell function assay in pediatric heart transplant recipients: Pediatr Transplant, 2014; 18(5); 485-90
10. Horwitz PA, Tsai EJ, Putt ME, Detection of cardiac allograft rejection and response to immunosuppressive therapy with peripheral blood gene expression: Circulation, 2004; 110(25); 3815-21
11. Moayedi Y, Foroutan F, Miller R, Risk evaluation using gene expression screening to monitor for acute cellular rejection in heart transplant recipients: J Heart Lung Transplant, 2019; 38(1); 51-58
12. Granata S, Santoro G, Signorini L, Comparative transcriptome analysis of peripheral blood mononuclear cells in renal transplant recipients in everolimus- and tacrolimus-based immunosuppressive therapy: Eur J Pharmacol, 2019; 859; 172494
13. Barwick BG, Neri P, Bahlis NJ, Multiple myeloma immunoglobulin lambda translocations portend poor prognosis: Nat Commun, 2019; 10(1); 1911
14. Binder C, Cvetkovski F, Sellberg F, CD2 Immunobiology: Front Immunol, 2020; 11; 1090
15. Mohs A, Kuttkat N, Reissing J, Functional role of CCL5/RANTES for HCC progression during chronic liver disease: J Hepatol, 2017; 66(4); 743-53
16. Hammoutene A, Biquard L, Lasselin J, A defect in endothelial autophagy occurs in patients with non-alcoholic steatohepatitis and promotes inflammation and fibrosis: J Hepatol, 2020; 72(3); 528-38
17. Adam N, Coutance G, Viailly PJ, Reverse transcriptase multiplex ligation-dependent probe amplification in endomyocardial biopsies for the diagnosis of cardiac allograft rejection: J Heart Lung Transplant, 2020; 39(2); 115-24
18. Kawamoto A, Nagata S, Anzai S, Ubiquitin D is upregulated by synergy of notch signalling and TNF-alpha in the inflamed intestinal epithelia of IBD patients: J Crohns Colitis, 2019; 13(4); 495-509
19. Yamaguchi T, Kitaya K, Daikoku N, Potential selectin L ligands involved in selective recruitment of peripheral blood CD16(-) natural killer cells into human endometrium: Biol Reprod, 2006; 74(1); 35-40
20. Park HJ, Kim BG, Lee SJ, Proteomic profiling of endothelial cells in human lung cancer: J Proteome Res, 2008; 7(3); 1138-50
21. Montecucco F, Braunersreuther V, Lenglet S, CC chemokine CCL5 plays a central role impacting infarct size and post-infarction heart failure in mice: Eur Heart J, 2012; 33(15); 1964-74
22. Lee YL, Ito K, Pi WC, Mediator subunit MED1 is required for E2A-PBX1-mediated oncogenic transcription and leukemic cell growth: Proc Natl Acad Sci USA, 2021; 118(6); e1922864118
23. Papac-Milicevic N, Breuss JM, Zaujec J, The interferon stimulated gene 12 inactivates vasculoprotective functions of NR4A nuclear receptors: Circ Res, 2012; 110(8); e50-63
24. Wang JH, Wei ZF, Gao YL, Activation of the mammalian target of rapamycin signaling pathway underlies a novel inhibitory role of ring finger protein 182 in ventricular remodeling after myocardial ischemia-reperfusion injury: J Cell Biochem, 2018 [Online ahead of print]
25. Liu YL, Ang SO, Weigent DA, Regulation of ferrochelatase gene expression by hypoxia: Life Sci, 2004; 75(17); 2035-43
26. Lim M, Brown HM, Kind KL, Haemoglobin expression in in vivo murine preimplantation embryos suggests a role in oxygen-regulated gene expression: Reprod Fertil Dev, 2019; 31(4); 724-34
27. Pu JJ, Redman CM, Visser JW, Lee S, Onset of expression of the components of the Kell blood group complex: Transfusion, 2005; 45(6); 969-74
28. Li CY, Zhan YQ, Xu CW, EDAG regulates the proliferation and differentiation of hematopoietic cells and resists cell apoptosis through the activation of nuclear factor-kappa B: Cell Death Differ, 2004; 11(12); 1299-308
29. Liu C, Liu B, Liu L, Arachidonic acid metabolism pathway is not only dominant in metabolic modulation but associated with phenotypic variation after acute hypoxia exposure: Front Physiol, 2018; 9; 236
30. Skov V, Burton M, Thomassen M, A 7-gene signature depicts the biochemical profile of early prefibrotic myelofibrosis: PLoS One, 2016; 11(8); e161570
31. Foley RN, Emerging erythropoiesis-stimulating agents: Nat Rev Nephrol, 2010; 6(4); 218-23
32. Shen Z, Gong W, Identification of candidate biomarkers in peripheral blood for cardiac allograft rejection based on bioinformatics analysis: Ann Transplant, 2015; 20; 312-19
33. Szeto C, Bloom JI, Sloane H, Impact of HLA-DR antigen binding cleft rigidity on T cell recognition: Int J Mol Sci, 2020; 21(19); 7081
34. Zhu S, Liu K, Chai Y, Divergent peptide presentations of HLA-A(*)30 alleles revealed by structures with pathogen peptides: Front Immunol, 2019; 10; 1709
35. Bunsuz A, Sercinoglu O, Ozbek P, Computational investigation of peptide binding stabilities of HLA-B*27 and HLA-B*44 alleles: Comput Biol Chem, 2020; 84; 107195
36. Horowitz A, Djaoud Z, Nemat-Gorgani N, Class I HLA haplotypes form two schools that educate NK cells in different ways: Sci Immunol, 2016; 1(3); eaag1672
37. Shboul M, Roschger P, Ganger R, Bone matrix hypermineralization associated with low bone turnover in a case of Nasu-Hakola disease: Bone, 2019; 123; 48-55
38. Kalaora S, Lee JS, Barnea E, Immunoproteasome expression is associated with better prognosis and response to checkpoint therapies in melanoma: Nat Commun, 2020; 11(1); 896
39. Min SH, Suzuki A, Weaver L, PIKfyve deficiency in myeloid cells impairs lysosomal homeostasis in macrophages and promotes systemic inflammation in mice: Mol Cell Biol, 2019; 39(21); e00158-19
40. Segarra-Mondejar M, Casellas-Diaz S, Ramiro-Pareta M, Synaptic activity-induced glycolysis facilitates membrane lipid provision and neurite outgrowth: EMBO J, 2018; 37(9); e97368
41. Chen Z, Liu J, Chen Q, Down-regulation of UBA6 exacerbates brain injury by inhibiting the activation of Notch signaling pathway to promote cerebral cell apoptosis in rat acute cerebral infarction model: Mol Cell Probes, 2020; 53; 101612
42. Gallagher PG, Disorders of erythrocyte hydration: Blood, 2017; 130(25); 2699-708
43. Ko WY, Kaercher KA, Giombini E, Effects of natural selection and gene conversion on the evolution of human glycophorins coding for MNS blood polymorphisms in malaria-endemic African populations: Am J Hum Genet, 2011; 88(6); 741-54
44. Liu J, Li Y, Tong J, Long non-coding RNA-dependent mechanism to regulate heme biosynthesis and erythrocyte development: Nat Commun, 2018; 9(1); 4386
45. Yu X, Kong Y, Dore LC, An erythroid chaperone that facilitates folding of alpha-globin subunits for hemoglobin synthesis: J Clin Invest, 2007; 117(7); 1856-65
46. Lechauve C, Butcher JT, Freiwan A, Endothelial cell alpha-globin and its molecular chaperone alpha-hemoglobin-stabilizing protein regulate arteriolar contractility: J Clin Invest, 2018; 128(11); 5073-82
47. Ogawa T, de Bold AJThe endocrine heart and inflammation: Medicina (B Aires), 2013; 73(6); 562-66 [in Spanish]
48. Talha S, Charloux A, Piquard F, Geny B, Brain natriuretic peptide and right heart dysfunction after heart transplantation: Clin Transplant, 2017; 31(6); ctr.12969
49. Hollander Z, Lin D, Chen V, Whole blood biomarkers of acute cardiac allograft rejection: double-crossing the biopsy: Transplantation, 2010; 90(12); 1388-93
Figures
 Figure 1. Differential expression of genes from endomyocardial biopsy and peripheral blood samples(A) Volcano plot of 57 502 genes expressed differentially in endomyocardial biopsy of patients with and without cardiac rejection. (B) Volcano plot of 131 624 genes expressed differentially in peripheral blood of patients with and without cardiac rejection. Red plots represent significantly upregulated genes (logFC ≥1, P value <0.05), blue plots are significantly downregulated genes (logFC ≤−1, P value <0.05), while the gray ones demonstrate genes with non-differential expression. Volcano plot graphs were created using the OmicStudio tools available at https://www.omicstudio.cn/tool.
Figure 1. Differential expression of genes from endomyocardial biopsy and peripheral blood samples(A) Volcano plot of 57 502 genes expressed differentially in endomyocardial biopsy of patients with and without cardiac rejection. (B) Volcano plot of 131 624 genes expressed differentially in peripheral blood of patients with and without cardiac rejection. Red plots represent significantly upregulated genes (logFC ≥1, P value <0.05), blue plots are significantly downregulated genes (logFC ≤−1, P value <0.05), while the gray ones demonstrate genes with non-differential expression. Volcano plot graphs were created using the OmicStudio tools available at https://www.omicstudio.cn/tool. Figure 2. PPI Networks of significant DEGs in endomyocardial biopsy and peripheral blood sample constructed with Cytoscape 3.8.2 software(A) Significant DEGs PPI networks of myocardium. (B) Significant DEGs PPI networks of peripheral blood. Nodes represent genes. Edges represent the interaction among proteins. the structure of proteins can be checked in nodes. Colors of edges represent the interaction between the proteins proved by different kinds of evidence. PPI – protein–protein interaction.
Figure 2. PPI Networks of significant DEGs in endomyocardial biopsy and peripheral blood sample constructed with Cytoscape 3.8.2 software(A) Significant DEGs PPI networks of myocardium. (B) Significant DEGs PPI networks of peripheral blood. Nodes represent genes. Edges represent the interaction among proteins. the structure of proteins can be checked in nodes. Colors of edges represent the interaction between the proteins proved by different kinds of evidence. PPI – protein–protein interaction. Figure 3. Networks of top 10 hub genes with connective degree of ≥10 achieved with CytoHubba, a plugin in Cytoscape(A) Ten hub genes of DEGs from endomyocardial biopsy samples. (B) Ten hub genes of DEGs from peripheral blood. The depth of color of each gene reflects its degree of connection.
Figure 3. Networks of top 10 hub genes with connective degree of ≥10 achieved with CytoHubba, a plugin in Cytoscape(A) Ten hub genes of DEGs from endomyocardial biopsy samples. (B) Ten hub genes of DEGs from peripheral blood. The depth of color of each gene reflects its degree of connection. Tables
 Table 1. Top 10 differentially expressed genes (DEGs) from endomyocardial biopsy samples.
Table 1. Top 10 differentially expressed genes (DEGs) from endomyocardial biopsy samples. Table 2. Top 10 differentially expressed genes (DEGs) from peripheral blood samples.
Table 2. Top 10 differentially expressed genes (DEGs) from peripheral blood samples. Table 3. Enrichment of Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) analysis of significance for differentially expressed genes (DEGs) from endomyocardial biopsy samples.
Table 3. Enrichment of Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) analysis of significance for differentially expressed genes (DEGs) from endomyocardial biopsy samples. Table 4. Enrichment of Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) analysis of significance for differentially expressed genes (DEGs) from peripheral blood samples.
Table 4. Enrichment of Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) analysis of significance for differentially expressed genes (DEGs) from peripheral blood samples. Table 5. Top 10 hub genes from endomyocardial biopsy samples enriched in Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG).
Table 5. Top 10 hub genes from endomyocardial biopsy samples enriched in Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG). Table 6. Top 10 hub genes from peripheral blood samples enriched in Gene Ontology (GO).
Table 6. Top 10 hub genes from peripheral blood samples enriched in Gene Ontology (GO). Table 1. Top 10 differentially expressed genes (DEGs) from endomyocardial biopsy samples.
Table 1. Top 10 differentially expressed genes (DEGs) from endomyocardial biopsy samples. Table 2. Top 10 differentially expressed genes (DEGs) from peripheral blood samples.
Table 2. Top 10 differentially expressed genes (DEGs) from peripheral blood samples. Table 3. Enrichment of Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) analysis of significance for differentially expressed genes (DEGs) from endomyocardial biopsy samples.
Table 3. Enrichment of Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) analysis of significance for differentially expressed genes (DEGs) from endomyocardial biopsy samples. Table 4. Enrichment of Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) analysis of significance for differentially expressed genes (DEGs) from peripheral blood samples.
Table 4. Enrichment of Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) analysis of significance for differentially expressed genes (DEGs) from peripheral blood samples. Table 5. Top 10 hub genes from endomyocardial biopsy samples enriched in Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG).
Table 5. Top 10 hub genes from endomyocardial biopsy samples enriched in Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG). Table 6. Top 10 hub genes from peripheral blood samples enriched in Gene Ontology (GO).
Table 6. Top 10 hub genes from peripheral blood samples enriched in Gene Ontology (GO). In Press
Original article
Diagnostic Utility of FAR1 Methylation Levels in Hepatocellular Carcinoma Patients Undergoing Liver Transpl...Ann Transplant In Press; DOI: 10.12659/AOT.951568
Original article
Inferior Long-Term Outcome of Fatty Liver Allografts After Orthotopic Liver TransplantationAnn Transplant In Press; DOI: 10.12659/AOT.950589
Database Analysis
Identification and Validation of Liver Transplantation-Induced Acute Lung Injury Biomarkers Using a Bioinfo...Ann Transplant In Press; DOI: 10.12659/AOT.950289
Original article
Survival and Recurrence in Liver Transplant Patients With Intrahepatic Cholangiocarcinoma and Hepatocellula...Ann Transplant In Press; DOI: 10.12659/AOT.950997
Most Viewed Current Articles
24 Aug 2021 : Review article 18,372
Normothermic Machine Perfusion (NMP) of the Liver – Current Status and Future PerspectivesDOI :10.12659/AOT.931664
Ann Transplant 2021; 26:e931664
05 Apr 2022 : Original article 14,731
Impact of Statins on Hepatocellular Carcinoma Recurrence After Living-Donor Liver TransplantationDOI :10.12659/AOT.935604
Ann Transplant 2022; 27:e935604
22 Nov 2022 : Original article 14,244
Long-Term Effects of Everolimus-Facilitated Tacrolimus Reduction in Living-Donor Liver Transplant Recipient...DOI :10.12659/AOT.937988
Ann Transplant 2022; 27:e937988
29 Dec 2021 : Original article 13,752
Efficacy and Safety of Tacrolimus-Based Maintenance Regimens in De Novo Kidney Transplant Recipients: A Sys...DOI :10.12659/AOT.933588
Ann Transplant 2021; 26:e933588








