01 December 2021: Original Paper
Solid-Phase C1q/C3d Fixing Readouts Correlate with High Median Fluorescence Intensity (MFI) De Novo Donor-Specific HLA Antibodies and C4d Antibody-Mediated Rejection in Kidney Transplant Recipients
Vasishta S. Tatapudi1BDEF, Dessislava Kopchaliiska2ABC, Gilberto J. da Gente2C, Owen F. Buenaventura2BC, Manpreet Singh3BC, Zoltan Laszik4BC, Deborah B. Adey1CDE, Raja RajalingamDOI: 10.12659/AOT.934175
Ann Transplant 2021; 26:e934175
Abstract
BACKGROUND: Solid-phase assays to investigate the complement-activating capacity of HLA antibodies have been utilized to optimize organ allocation and improve transplant outcomes. The clinical utility of C1q/C3d-binding characteristics of de novo donor-specific anti-HLA antibodies (dnDSA) associated with C4d-positive antibody-mediated rejection (C4d⁺ AMR) in kidney transplants (KTx) has not been defined.
MATERIAL AND METHODS: Sera from 120 KTx recipients that had dnDSA concurrent with protocol/cause biopsy (median 3.8 years after transplantation) were screened for C1q and C3d-binding dnDSA. The difference in the incidence of C4d⁺ AMR between recipients with and without C1q/C3d-binding dnDSA was assessed.
RESULTS: Over 86% of dnDSAs were class II antibodies. The immunodominant dnDSAs characterized by the highest median fluorescence intensity (MFI) in most recipients were HLA-DQ antibodies (67%). Most recipients (62%, n=74) had either C1q⁺ (56%), C3d⁺ (48%), or both C1q⁺C3d⁺ (41.2%) dnDSA, while the remaining 38% were negative for both C1q and C3d. Of those with C1q⁺/C3d⁺ dnDSA, 87% had high-MFI IgG (MFI=14144±5363 and 13932±5278, respectively), while 65% of C1q⁻C3d⁻ dnDSA had low-MFI IgG (MFI=5970±3347). The incidence of C4d+ AMR was significantly higher in recipients with C1q⁺ (66%), C3d+ (74%), and C1q⁺C3d⁺ (72%) dnDSA than in those with C1q⁻C3d⁻ dnDSA (30%) recipients. Recipients with C3d⁺/C1q⁺ dnDSA had higher C4d⁺ scores on biopsy.
CONCLUSIONS: C1q⁺/C3d⁺ dnDSA were associated with C4d⁺ AMR and high-IgG MFI. Our data call into question the predictive utility of C1q/C3d-binding assays in identifying KTx recipients at risk of allograft failure. In conclusion, IgG MFI is sufficient for clinical management, and the C1q/C3d-assays with added cost do not provide any additional information.
Keywords: HLA Antigens, Kidney Transplantation, Organ Transplantation, Complement C1q, Graft Rejection, Humans, Isoantibodies, transplant recipients
Background
Patel and Terasaki’s seminal study in 1969 established the role of donor-specific anti-HLA antibodies in the pathogenesis of hyper-acute kidney transplant rejection [1]. Kidney transplantation is considered the treatment of choice for end-stage renal disease (ESRD), given its positive impact on patient survival and quality of life relative to dialytic therapies. This is a testament to the advances in our insights into the mechanistic underpinnings of alloimmunity and the development of potent immunosuppressive therapies [2,3]. Since the advent of calcineurin inhibitors (CNIs) in the 1980s, acute rejection rates have declined considerably [4]. However, this has not resulted in improved long-term graft survival [5]. Antibody-mediated rejection (AMR) is the primary driver of renal allograft failure and is responsible for this discordance between long-term graft survival rates despite lower rates of acute rejection [6]. AMR occurs when donor-specific antibodies (DSA) interact with mismatched human leukocyte antigens (HLA), resulting in complement-mediated endothelial injury [2,7]. C4d is a complement protein that is conspicuously deposited in the renal peritubular capillaries during antibody-mediated rejection [8]. Positive C4d staining in the peritubular capillaries of kidney transplant biopsies is a marker of HLA binding by DSA and consequent complement activation [9]. The advent of solid-phase Luminex platform-based assays for HLA antibody detection has enhanced our ability to diagnose AMR [10,11]. In 2003, C4d positivity on histology and circulating DSA were incorporated in the Banff classification system as diagnostic criteria for AMR [12].
However, there is an emerging consensus that all DSAs are not created equal in terms of their pathogenic potential and that DSAs that bind complement are associated with AMR and graft loss [13]. Loupy et al reported on the association between C1q binding DSAs and the risk of renal allograft failure. Patients with detectable complement-binding DSAs after transplant had inferior 5-year renal allograft survival than those with non-complement-binding DSAs and patients with no DSA [14]. C3d is another complement split product that results from the cleavage of C3 by C3-convertases. Detection of C3d-binding DSA at the time of AMR diagnosis is predictive of renal graft loss [7]. A better comprehension of the etiopathogenesis of AMR and the critical contribution of the complement system supports the role of targeted complement blockade in its treatment [3]. Terminal complement inhibitors, eculizumab, and C1 esterase inhibitor (C1-INH) have been tested in clinical trials [15–17]. However, the clinical utility of C1q/C3d-binding characteristics of de novo donor-specific anti-HLA antibodies (dnDSA) associated with C4d-positive antibody-mediated rejection (C4d+ AMR) in kidney transplants (KTx) has not been defined. Risk stratification of patients based on these assays may allow us to personalize anti-rejection therapy and minimize their adverse effects. However, some have argued that the C1q-binding activity of DSA may be a correlation of antibody strength and that it does not necessarily signify a qualitative difference in pathogenic potential [18]. In the present study, we determined the nature of dnDSA, their capacity to fix complement components (C1q and C3d) in solid-phase assays, and their correlation with biopsy-proven C4d+ AMR in kidney transplant recipients. In addition, we measured the differences between C1q+/C3d+ and C1q−C3d− in dnDSA MFI to assess the clinical utility of complement-binding assays.
Material and Methods
STUDY COHORT:
All kidney, kidney and pancreas transplant recipients at the University of California San Francisco (UCSF) Medical Center, who underwent a 6-month protocol or indication for renal allograft biopsy between 01/01/2009 and 06/30/2012, were assessed. The Institutional Review Board of the UCSF approved this retrospective study (IRB# 14-19103).
HLA TYPING AND ANTIBODY TESTING:
Serum samples collected at the time of biopsy were tested for HLA antibodies using Luminex-based LABScreenTM single antigen bead assays according to the manufacturer’s instructions (One Lambda, Canoga Park, CA). Serum samples were pre-treated with dithiothreitol (DTT) to prevent aggregation of high titer antibodies (prozone effect) and to increase the sensitivity for IgG antibody detection [19,20]. The median fluorescence intensity (MFI) adjusted for background negative control bead reactivity was used as an arbitrary measure of antibody quantity. All antibody testing was performed using the LABXpress™ Pipettor, high throughput automation that aspirates and dispenses precise volumes into test wells of a 96-well reaction plate and reagent vials (One Lambda, Canoga Park, CA) to minimize inter-assay variations. Calculated Panel Reactive Antibody (CPRA) values were determined using UNOS CPRA calculator by entering the targets (ie, unacceptable antigens) of identified HLA antibodies using the following criteria: HLA-A/B/C/DRB1/DRB3/DRB4/DRB5/DQB1 allotypes are listed as unacceptable antigens if patient displays antibodies with ≥2,000 MFI reactivity against these allotypes; Bw4 or Bw6 are listed as unacceptable antigens if patient displays antibodies to these epitopes at any MFI; HLA-DQA1, DPA1 and DPB1 specificities of any MFI are not listed as unacceptable antigens.
A cohort of 120 kidney transplant recipients who had detectable dnDSA at the time of biopsy (timing between serum assessed for dnDSA and biopsy is ±7 days) were included in this study. Only those DSA that were not detectable in the pre-transplant sera (tested quarterly for over a year) but were detectable in post-transplant sera were designated as dnDSA. If more than one dnDSA was detected, dnDSA with the highest MFI was considered as an immunodominant dnDSA. Renal allograft pathology findings were reviewed from recipient medical records, and diagnoses were made according to the Banff classification by clinical pathologists. Intermediate level typing of all 11 HLA genes (HLA-A, B, C, DRB1, DRB3, DRB4, DRB5, DQA1, DQB1, DPA1, and DPB1) was performed for all donors and recipients using Luminex-based LABType SSO typing (One Lambda, Canoga Park, CA, USA) according to the manufacturer’s protocol.
Serum samples displaying dnDSA at the time of biopsy were also tested for C1q-binding HLA antibodies using C1qScreen according to the manufacturer’s instructions (One Lambda, Canoga Park, CA). Serum samples were further tested for HLA antibodies (DTT-treated serum) and C3d-binding HLA antibodies (DTT untreated serum) using the LIFECODES® single antigen bead assay with and without the C3d detection system (Immucor, Norcross, GA). All antibody tests were independently analyzed in a blinded manner.
HISTOLOGY AND IMMUNOCHEMISTRY ANALYSES:
Renal biopsies were fixed in AFA (acetic acid-formol-absolute alcohol) fixative and stained by standard methods for routine microscopy. The biopsies were reviewed by 2 renal pathologists and a nephrologist who was blinded to the clinical information. Histological changes were graded according to Banff classification [21]. C4d staining was performed on paraffin sections using human polyclonal antiserum against C4d (Biomedica Gruppe, Austria). The C4d score varied from 0 to 3 (negative, minimal, focal, and diffuse) depending on the percentage of peritubular capillary (PTC) with a linear staining pattern: no staining=score 0, <10%=score 1 (minimal), 10–50%=score 2 (focal), and >50%=score 3 (diffuse) [22]. A diagnosis of acute cellular or humoral rejection was based on published criteria in conjunction with allograft dysfunction [12]. Humoral rejection was defined based on Banff criteria classification [21].
STATISTICAL ANALYSIS:
In the data analysis, continuous variables were expressed as mean±SD and compared using two-tailed Fisher’s exact probabilities. Statistical significance was set at
Results
BASELINE CHARACTERISTICS OF THE STUDY POPULATION:
The study cohort comprised 120 kidney transplant recipients who developed dnDSA at the time of biopsy. Biopsies were performed at a median duration of 3.8 years after transplant. Demographic and clinical characteristics are provided in Table 1. The mean age of the recipients was 47±18 years (mean±SD). Forty% of the recipients were women. The majority (86%) were primary transplant recipients (first transplants). Living donor transplant recipients comprised 30%, and deceased donor transplant recipients constituted the remainder. Kidney and pancreas transplant recipients accounted for 9% of the patients. The mean number of HLA mismatches detectable was 7.7±2.3 at the HLA-A, -B, -C, -DR, and -DQ loci, respectively.
C1Q AND C3D BINDING CHARACTERISTICS OF DNDSA:
A majority of the kidney transplant recipients had dnDSA to a single mismatched donor HLA allotype (53.4%), 23.3% had dnDSA to 2 HLA allotypes, and 23.3% had dnDSA to >2 mismatched donor HLA allotypes. dnDSA was directed against HLA class II antigens in most recipients (86%, 103/120). The highest MFI dnDSA (also called immunodominant dnDSA) in most recipients was directed to HLA-DQ antigens (67%, 80/120). A total of 62% (n=74) of the recipients had C1q+ and/or C3d+ dnDSA [C3d+C1q−=7, C3d+C1q+=50, C3d−C1q+=17], 56% (n=67) had C1q+ dnDSA, and 48% (n=57) had C3d+ dnDSA (Figure 1). In the remaining 46 recipients (38%), dnDSAs did not bind to either C1q or C3d (i.e., C1q−C3d− dnDSA) (Figure 1). Most C1q+ or C3d+ dnDSA (86%, 103/120) were directed against HLA class II antigens (Figure 2). In particular, the majority of C1q+ or C3d+ dnDSA (66.7%, 80/120) were directed to HLA-DQ antigens (Figure 3).
:
More than half (53%, 63/120) of the recipients with dnDSA were diagnosed with C4d+ AMR (Table 2). The incidence of C4d+ AMR was more frequent in the C1q+ dnDSA group than in the C1q− dnDSA group (66% vs 34%, P=0.0017). Similarly, C4d+ AMR was more frequent in the C3d+ dnDSA group than in the C3d− dnDSA group (74% vs 33%, P=0.0001). Moreover, C4d+ AMR was more frequent in recipients with dnDSA with an IgG MFI ≥5000 compared to those with dnDSA of <5000 MFI (70% vs 36%, P<0.0003).
IGG MFI CORRELATES WITH C1Q/C3D BINDING CAPACITY:
When immunodominant dnDSA with the highest MFI was considered in each recipient, the C1q+ or C3d+ dnDSA groups had higher MFI (14144±5363 and 13932±5278, respectively) compared with the C3d−C1q− dnDSA (5970±3347) (Figure 1). The difference in mean MFI between C1q+ dnDSA vs C1q−C3d− dnDSA groups and C3d+ dnDSA vs C1q−C3d− dnDSA groups, both achieved statistical significance (P<0.0001). There was no statistically significant difference in mean MFI between the C1q+ dnDSA and C3d+ dnDSA groups (p=0.95). Sixteen kidney transplant recipients had dnDSA with an IgG MFI ≥5000 but negative for both C1q and C3d; 4 of them exhibited C4d+ AMR, and 2 showed C4d− AMR (Table 3). These 16 sera were re-tested for C1q and C3d after 1: 16 dilution, and results remained to be negative for both C1q and C3d-binding dnDSA.
C1Q/C3D BINDING CAPACITY AND CLINICAL OUTCOMES:
The group with C1q+/C3d+ dnDSA had a significantly higher proportion of recipients diagnosed with C4d+ AMR, acute AMR (aAMR), chronic AMR (cAMR), and acute cellular rejection (ACR) than the C1q−C3d− dnDSA group [AMR, 34/47 (47%) vs 14/46 (30%), p<0.0001; aAMR, 24/47 (51%) vs 6/46 (13%), p<0.0001; cAMR 18/47 (38%) vs 10/46 (22%), P=0.02; ACR 12/47 (26%) vs 6/46 (13%), P=0.03]. Moreover, patients with C1q+/C3d+ dnDSA had higher C4d scores on renal allograft biopsy (Figure 4).
Discussion
Antibody-mediated rejection is recognized as the dominant etiology of renal allograft failure, and its prevention and treatment are key to improving long-term graft outcomes [3,6]. In AMR, DSAs bind to mismatched HLAs expressed on the graft vascular endothelium. This is followed by the initiation of the classical complement cascade and the resultant allograft injury by the membrane attack complex and infiltrating mononuclear cells [23]. Detection of inert complement split products, C4d in the microcirculation, is believed to be a marker of DSA binding to HLAs and is a key diagnostic feature for AMR [3,9]. Recent studies point to the association between complement-binding ability of DSA and poor graft outcomes [7,14]. These studies espoused the role of C1q and C3d binding assays in recognizing kidney transplant recipients at risk of AMR and resultant graft loss. Others have argued that this is a reflection of DSA strength rather than a qualitative difference in pathogenicity. Yell et al suggested that C1q binding DSAs are not qualitatively distinct antibodies with pathogenic potential, but that the assay merely captures antibodies with greater MFI strength [18].
Our study found that a high percentage of complement-binding dnDSA was endowed with both C3d and C1q binding capacity. This correlation is not unexpected because C1q binding sets in motion a cascade of events that include the formation of C3d. In the study by Sicard et al, C3d binding DSA identified concurrent with the diagnosis of AMR correlated with graft failure, and the presence of C1q binding DSAs showed a similar trend; this association was not statistically significant. Sicard et al surmise that the superior performance C3d binding assay relative to the C1q assays in predicting graft loss is a consequence of the greater sensitivity of the C3d-binding assay derived from the fact that each enzymatic reaction in the complement cascade greatly amplifies the generation of downstream elements [7]. Complement regulatory mechanisms that inhibit C3 convertase formation could limit the specificity of C1q-binding assays. Since detection of C3d signifies cleavage of C3, C3-binding DSAs may have a stronger association with AMR than C1q-binding DSAs. However, since longitudinal data were not collected as part of our study, we were unable to assess the impact of complement-binding antibodies on graft survival.
Data reported by Pollinger et al suggest that preexisting HLA class II DSAs confer risk for humoral rejection after transplantation [24]. In our cohort, a significant proportion (52.5%) of recipients with dnDSA were found to have C4d+AMR, and the high MFI dnDSAs in most recipients were directed to HLA-DQ antigens (class II). We believe that the high incidence of dnDSA, the nature of these antibodies (the majority being HLA class II antibodies), and the frequency of diagnosis of AMR in those with detectable dnDSAs justify post-transplant dnDSA screening protocols to identify at-risk patients. A case can be made for renal allograft biopsy and early therapeutic intervention in patients with rising titers of dnDSA (≥5000 MFI).
Conclusions
Our data indicated that MFI strength of dnDSAs provides clinically valuable information but does not support the utility of complement-binding assays in risk stratification of kidney transplant recipients. While C1q+/C3d+ dnDSA dnDSA was associated with C4d+ AMR, our data suggest that this effect reflects the correlation between complement-binding capacity and the MFI of dnDSA. This calls into question the predictive utility of C1q or C3d-binding solid-phase assays in identifying kidney transplant recipients at risk of allograft failure. Therapeutic interventions based on sophisticated knowledge of AMR immunopathogenesis, including plasmapheresis, IVIg, B-cell, and plasma-cell depleting agents, as well as pharmacologic complement blockade, must be employed judiciously to treat AMR if the inferior long-term graft outcomes are to be improved upon.
Figures
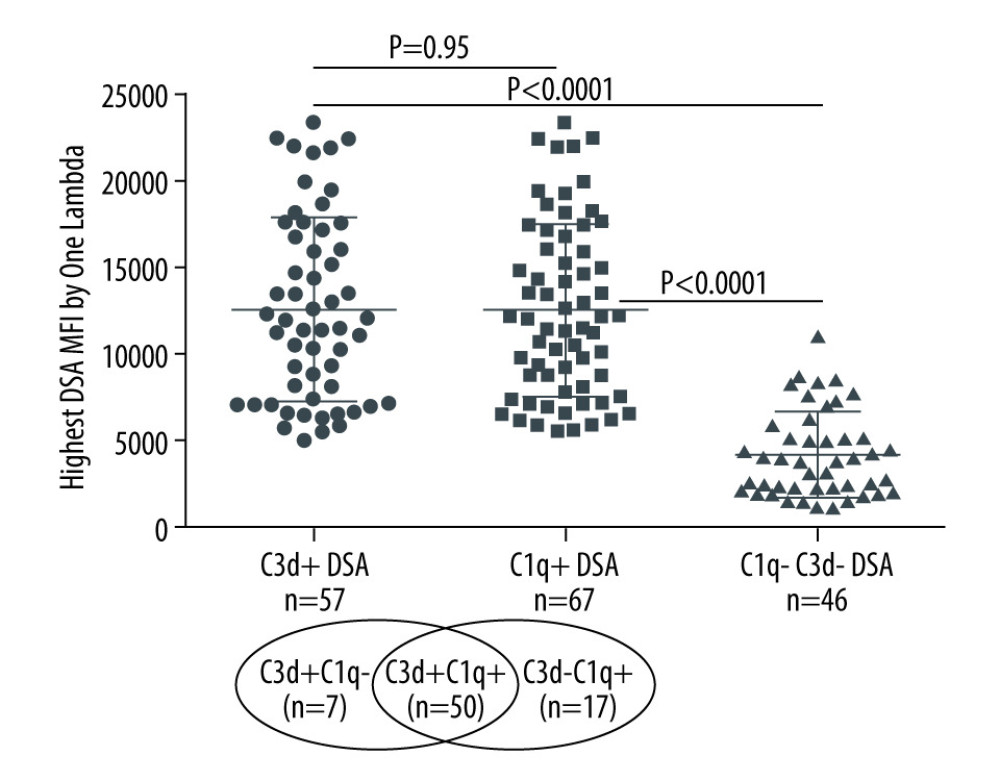 Figure 1. Solid-phase complement-binding characteristics of de novo donor-specific HLA antibodies (dnDSA) in kidney transplant recipientsRelationship between median fluorescence intensity (MFI) of de novo IgG DSA and ability of C1q or C3d-binding capacity. For recipients with multiple dnDSAs, only the immunodominant dnDSAs defined by the highest MFI are plotted. Some of the C3d+ DSA can be also positive for C1q, and vice versa.
Figure 1. Solid-phase complement-binding characteristics of de novo donor-specific HLA antibodies (dnDSA) in kidney transplant recipientsRelationship between median fluorescence intensity (MFI) of de novo IgG DSA and ability of C1q or C3d-binding capacity. For recipients with multiple dnDSAs, only the immunodominant dnDSAs defined by the highest MFI are plotted. Some of the C3d+ DSA can be also positive for C1q, and vice versa. 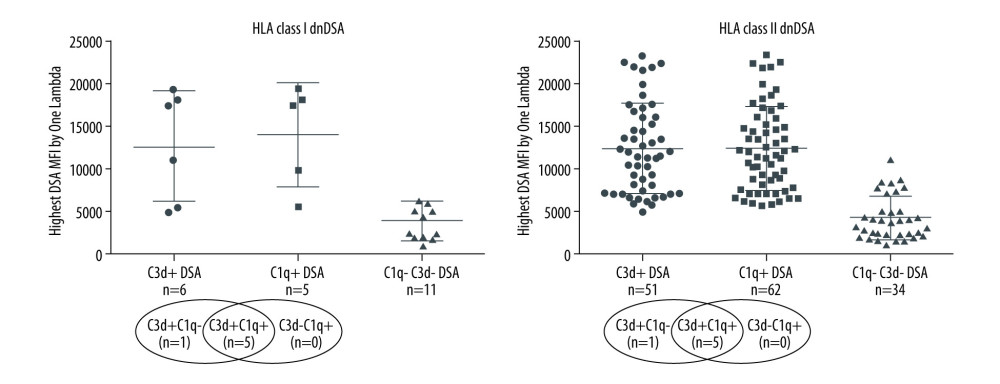 Figure 2. Nature of de novo donor-specific HLA antibodies (dnDSA) in kidney transplant recipientsMost de novo DSA in kidney transplant recipients are HLA class II antigen-specific. Shown is the relationship between median fluorescence intensity (MFI) of de novo class I and class II IgG DSA and ability of C1q or C3d-binding capacity. For recipients with multiple dnDSAs, only the immunodominant dnDSAs defined by the highest IgG MFI are plotted. Some of the C3d+ DSA can be also positive for C1q, and vice versa.
Figure 2. Nature of de novo donor-specific HLA antibodies (dnDSA) in kidney transplant recipientsMost de novo DSA in kidney transplant recipients are HLA class II antigen-specific. Shown is the relationship between median fluorescence intensity (MFI) of de novo class I and class II IgG DSA and ability of C1q or C3d-binding capacity. For recipients with multiple dnDSAs, only the immunodominant dnDSAs defined by the highest IgG MFI are plotted. Some of the C3d+ DSA can be also positive for C1q, and vice versa. 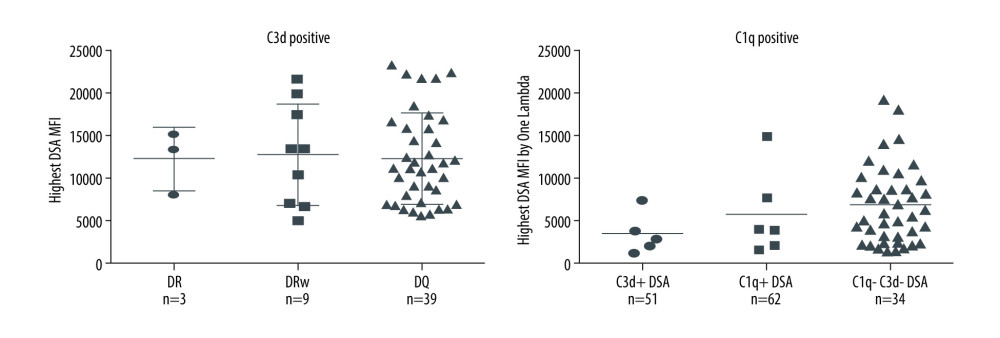 Figure 3. Nature of de novo donor-specific HLA class II antibodies in kidney transplant recipientsMost de novo DSA in kidney transplant recipients are HLA-DQ antigen-specific. Shown is the relationship between median fluorescence intensity (MFI) of de novo class II IgG DSA and the ability of C1q or C3d-binding capacity.
Figure 3. Nature of de novo donor-specific HLA class II antibodies in kidney transplant recipientsMost de novo DSA in kidney transplant recipients are HLA-DQ antigen-specific. Shown is the relationship between median fluorescence intensity (MFI) of de novo class II IgG DSA and the ability of C1q or C3d-binding capacity. 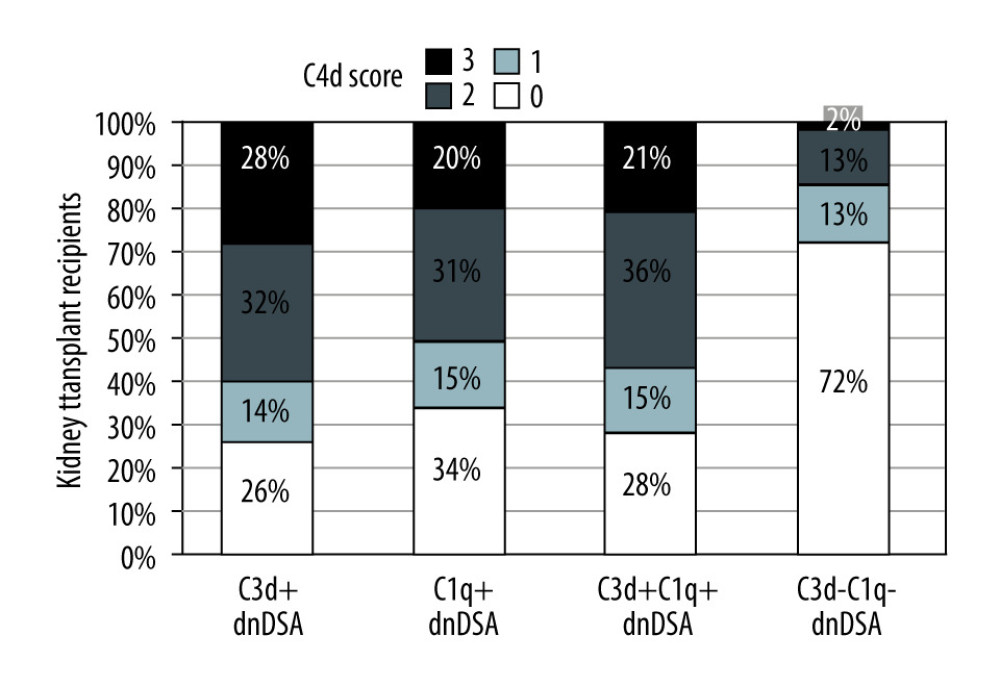 Figure 4. Correlation of C3d+ or C1q+ de novo donor-specific HLA class antibodies (dnDSA) with higher C4d scoresNo staining=score 0, <10%=score 1 (minimal), 10–50%=score 2 (focal) and >50%=score 3 (diffuse).
Figure 4. Correlation of C3d+ or C1q+ de novo donor-specific HLA class antibodies (dnDSA) with higher C4d scoresNo staining=score 0, <10%=score 1 (minimal), 10–50%=score 2 (focal) and >50%=score 3 (diffuse). Tables
Table 1. Demographic and clinical characteristics of kidney transplant recipients developing de novo donor-specific HLA antibodies (n=120).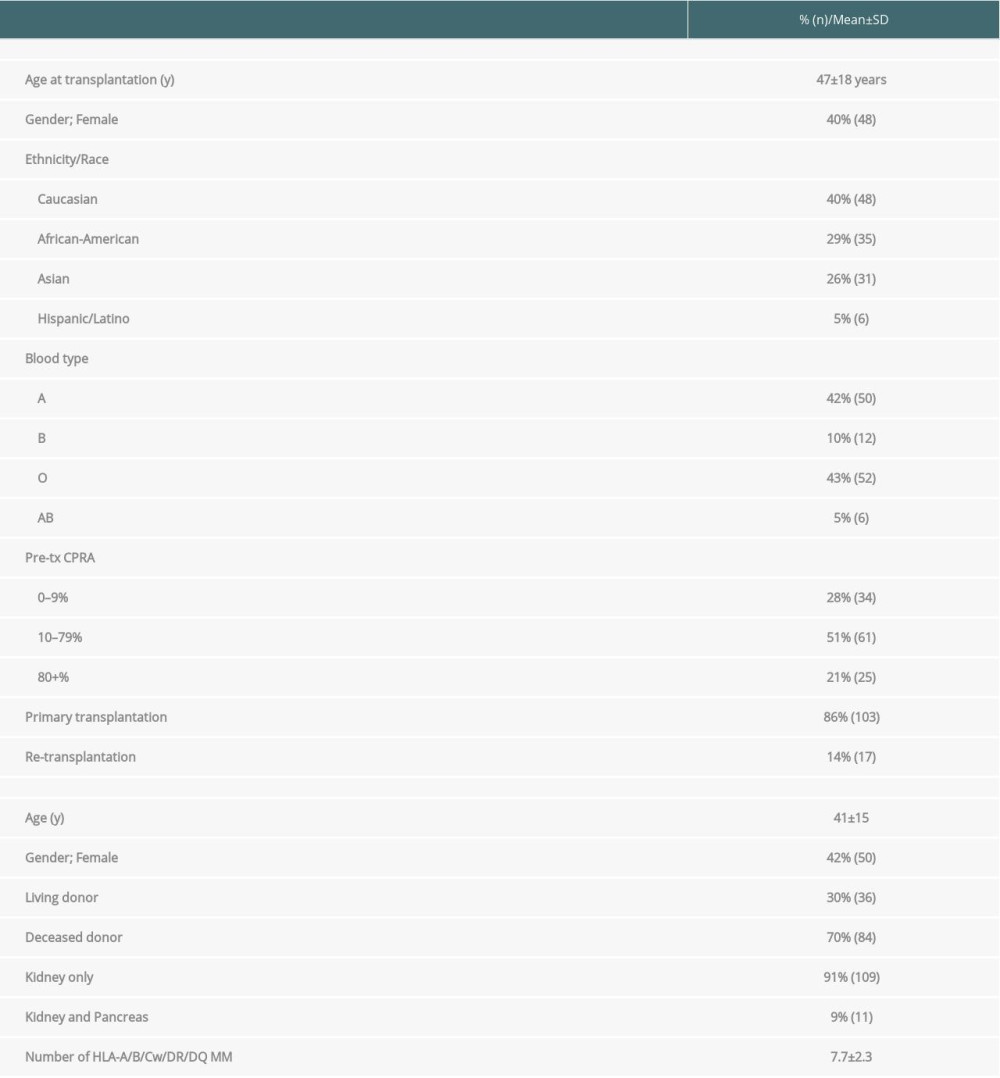 Table 2. The C1q-binding de novo donor-specific HLA antibodies (dnDSA), C3d-binding dnDSA, and IgG ≥5000 MFI dnDSA are associated with C4d+ antibody-mediated rejection (AMR).
Table 2. The C1q-binding de novo donor-specific HLA antibodies (dnDSA), C3d-binding dnDSA, and IgG ≥5000 MFI dnDSA are associated with C4d+ antibody-mediated rejection (AMR).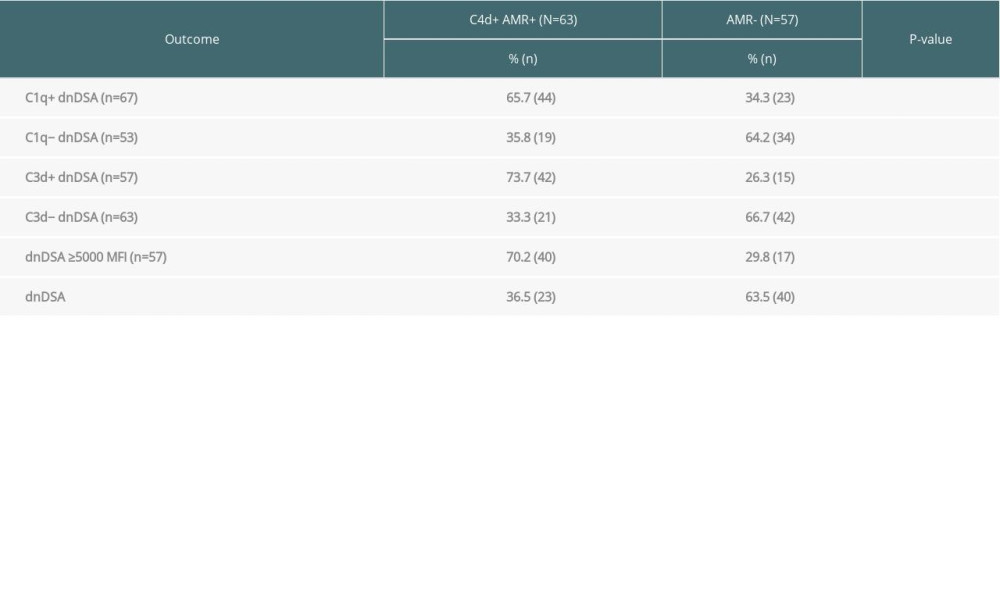 Table 3. Kidney transplant recipients with de novo donor-specific HLA antibodies (dnDSA) of IgG MFI ≥5000 but negative for C3d and C1q-binding. The median fluorescence intensity (MFI) of each dnDSA is given in parenthesis.
Table 3. Kidney transplant recipients with de novo donor-specific HLA antibodies (dnDSA) of IgG MFI ≥5000 but negative for C3d and C1q-binding. The median fluorescence intensity (MFI) of each dnDSA is given in parenthesis.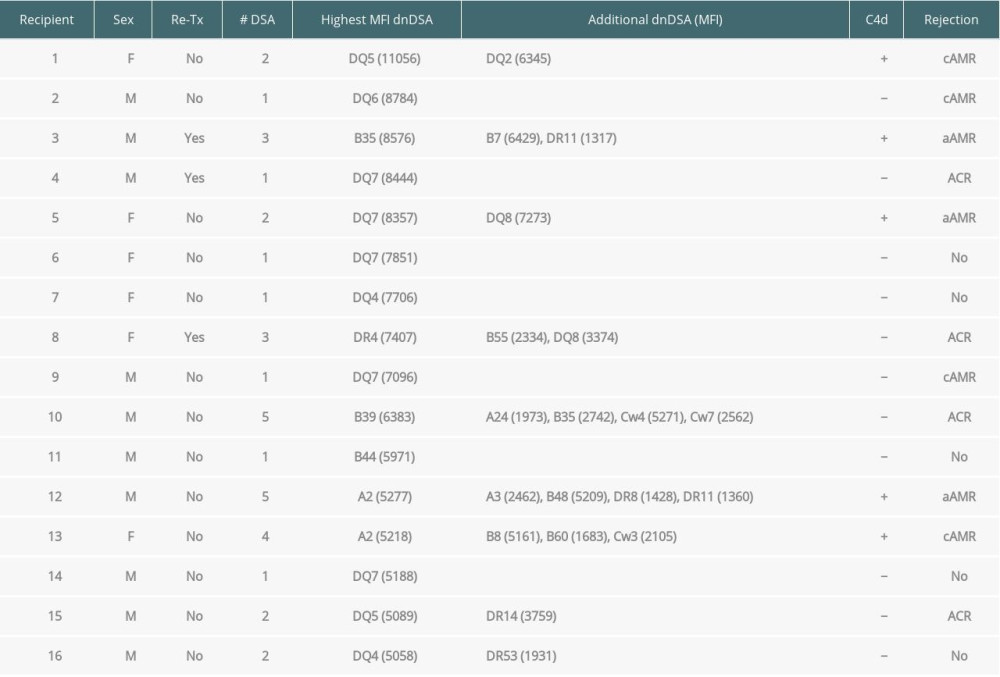
References
1. Patel R, Terasaki PI, Significance of the positive crossmatch test in kidney transplantation: N Engl J Med, 1969; 280; 735-39
2. Nankivell BJ, Alexander SI, Rejection of the kidney allograft: N Engl J Med, 2010; 363; 1451-62
3. Tatapudi VS, Montgomery RA, Pharmacologic complement inhibition in clinical transplantation: Curr Transplant Rep, 2017; 4; 91-100
4. Hariharan S, Johnson CP, Bresnahan BA, Improved graft survival after renal transplantation in the United States, 1988 to 1996: N Engl J Med, 2000; 342; 605-12
5. Meier-Kriesche HU, Schold JD, Lack of improvement in renal allograft survival despite a marked decrease in acute rejection rates over the most recent era: Am J Transplant, 2004; 4; 378-83
6. Gaston RS, Cecka JM, Kasiske BL, Evidence for antibody-mediated injury as a major determinant of late kidney allograft failure: Transplantation, 2010; 90; 68-74
7. Sicard A, Ducreux S, Rabeyrin M, Detection of C3d-binding donor-specific anti-HLA antibodies at diagnosis of humoral rejection predicts renal graft loss: J Am Soc Nephrol, 2015; 26; 457-67
8. Feucht HE, Felber E, Gokel MJ, Vascular deposition of complement-split products in kidney allografts with cell-mediated rejection: Clin Exp Immunol, 1991; 86; 464-70
9. Haas M, Sis B, Racusen LC, Banff 2013 meeting report: inclusion of c4d-negative antibody-mediated rejection and antibody-associated arterial lesions: Am J Transplant, 2014; 14; 272-83
10. Karpinski M, Rush D, Jeffery J, Flow cytometric crossmatching in primary renal transplant recipients with a negative anti-human globulin enhanced cytotoxicity crossmatch: J Am Soc Nephrol, 2001; 12; 2807-14
11. Tait BD, Hudson F, Cantwell L, Review article: Luminex technology for HLA antibody detection in organ transplantation: Nephrology (Carlton), 2009; 14; 247-54
12. Racusen LC, Colvin RB, Solez K, Antibody-mediated rejection criteria – an addition to the Banff 97 classification of renal allograft rejection: Am J Transplant, 2003; 3; 708-14
13. Loupy A, Hill GS, Jordan SC, The impact of donor-specific anti-HLA antibodies on late kidney allograft failure: Nat Rev Nephrol, 2012; 8; 348-57
14. Loupy A, Lefaucheur C, Vernerey D, Complement-binding anti-HLA antibodies and kidney-allograft survival: N Engl J Med, 2013; 369; 1215-26
15. Stegall MD, Diwan T, Raghavaiah S, Terminal complement inhibition decreases antibody-mediated rejection in sensitized renal transplant recipients: Am J Transplant, 2011; 11; 2405-13
16. Viglietti D, Gosset C, Loupy A, C1 Inhibitor in acute antibody-mediated rejection nonresponsive to conventional therapy in kidney transplant recipients: A pilot study: Am J Transplant, 2016; 16; 1596-603
17. Montgomery RA, Orandi BJ, Racusen L, Plasma-derived C1 esterase inhibitor for acute antibody-mediated rejection following kidney transplantation: Results of a randomized double-blind placebo-controlled pilot study: Am J Transplant, 2016; 16; 3468-78
18. Yell M, Muth BL, Kaufman DB, C1q binding activity of de novo donor-specific HLA antibodies in renal transplant recipients with and without antibody-mediated rejection: Transplantation, 2015; 99; 1151-55
19. Kosmoliaptsis V, Chaudhry AN, Sharples LD, Predicting HLA class I alloantigen immunogenicity from the number and physiochemical properties of amino acid polymorphisms: Transplantation, 2009; 88; 791-98
20. Weinstock C, Schnaidt M, The complement-mediated prozone effect in the Luminex single-antigen bead assay and its impact on HLA antibody determination in patient sera: Int J Immunogenet, 2013; 40; 171-77
21. Solez K, Colvin RB, Racusen LC, Banff ‘05 Meeting Report: Differential diagnosis of chronic allograft injury and elimination of chronic allograft nephropathy (‘CAN’): Am J Transplant, 2007; 7; 518-26
22. Solez K, Colvin RB, Racusen LC, Banff ‘07 classification of renal allograft pathology: Updates and future directions: Am J Transplant, 2008; 8; 753-60
23. Montgomery RA, Lonze BE, Tatapudi VS: Transplantation, 2018; 102; 2-4
24. Pollinger HS, Stegall MD, Gloor JM, Kidney transplantation in patients with antibodies against donor HLA class II: Am J Transplant, 2007; 7; 857-63
Figures
 Figure 1. Solid-phase complement-binding characteristics of de novo donor-specific HLA antibodies (dnDSA) in kidney transplant recipientsRelationship between median fluorescence intensity (MFI) of de novo IgG DSA and ability of C1q or C3d-binding capacity. For recipients with multiple dnDSAs, only the immunodominant dnDSAs defined by the highest MFI are plotted. Some of the C3d+ DSA can be also positive for C1q, and vice versa.
Figure 1. Solid-phase complement-binding characteristics of de novo donor-specific HLA antibodies (dnDSA) in kidney transplant recipientsRelationship between median fluorescence intensity (MFI) of de novo IgG DSA and ability of C1q or C3d-binding capacity. For recipients with multiple dnDSAs, only the immunodominant dnDSAs defined by the highest MFI are plotted. Some of the C3d+ DSA can be also positive for C1q, and vice versa. Figure 2. Nature of de novo donor-specific HLA antibodies (dnDSA) in kidney transplant recipientsMost de novo DSA in kidney transplant recipients are HLA class II antigen-specific. Shown is the relationship between median fluorescence intensity (MFI) of de novo class I and class II IgG DSA and ability of C1q or C3d-binding capacity. For recipients with multiple dnDSAs, only the immunodominant dnDSAs defined by the highest IgG MFI are plotted. Some of the C3d+ DSA can be also positive for C1q, and vice versa.
Figure 2. Nature of de novo donor-specific HLA antibodies (dnDSA) in kidney transplant recipientsMost de novo DSA in kidney transplant recipients are HLA class II antigen-specific. Shown is the relationship between median fluorescence intensity (MFI) of de novo class I and class II IgG DSA and ability of C1q or C3d-binding capacity. For recipients with multiple dnDSAs, only the immunodominant dnDSAs defined by the highest IgG MFI are plotted. Some of the C3d+ DSA can be also positive for C1q, and vice versa. Figure 3. Nature of de novo donor-specific HLA class II antibodies in kidney transplant recipientsMost de novo DSA in kidney transplant recipients are HLA-DQ antigen-specific. Shown is the relationship between median fluorescence intensity (MFI) of de novo class II IgG DSA and the ability of C1q or C3d-binding capacity.
Figure 3. Nature of de novo donor-specific HLA class II antibodies in kidney transplant recipientsMost de novo DSA in kidney transplant recipients are HLA-DQ antigen-specific. Shown is the relationship between median fluorescence intensity (MFI) of de novo class II IgG DSA and the ability of C1q or C3d-binding capacity. Figure 4. Correlation of C3d+ or C1q+ de novo donor-specific HLA class antibodies (dnDSA) with higher C4d scoresNo staining=score 0, <10%=score 1 (minimal), 10–50%=score 2 (focal) and >50%=score 3 (diffuse).
Figure 4. Correlation of C3d+ or C1q+ de novo donor-specific HLA class antibodies (dnDSA) with higher C4d scoresNo staining=score 0, <10%=score 1 (minimal), 10–50%=score 2 (focal) and >50%=score 3 (diffuse). Tables
 Table 1. Demographic and clinical characteristics of kidney transplant recipients developing de novo donor-specific HLA antibodies (n=120).
Table 1. Demographic and clinical characteristics of kidney transplant recipients developing de novo donor-specific HLA antibodies (n=120). Table 2. The C1q-binding de novo donor-specific HLA antibodies (dnDSA), C3d-binding dnDSA, and IgG ≥5000 MFI dnDSA are associated with C4d+ antibody-mediated rejection (AMR).
Table 2. The C1q-binding de novo donor-specific HLA antibodies (dnDSA), C3d-binding dnDSA, and IgG ≥5000 MFI dnDSA are associated with C4d+ antibody-mediated rejection (AMR). Table 3. Kidney transplant recipients with de novo donor-specific HLA antibodies (dnDSA) of IgG MFI ≥5000 but negative for C3d and C1q-binding. The median fluorescence intensity (MFI) of each dnDSA is given in parenthesis.
Table 3. Kidney transplant recipients with de novo donor-specific HLA antibodies (dnDSA) of IgG MFI ≥5000 but negative for C3d and C1q-binding. The median fluorescence intensity (MFI) of each dnDSA is given in parenthesis. Table 1. Demographic and clinical characteristics of kidney transplant recipients developing de novo donor-specific HLA antibodies (n=120).
Table 1. Demographic and clinical characteristics of kidney transplant recipients developing de novo donor-specific HLA antibodies (n=120). Table 2. The C1q-binding de novo donor-specific HLA antibodies (dnDSA), C3d-binding dnDSA, and IgG ≥5000 MFI dnDSA are associated with C4d+ antibody-mediated rejection (AMR).
Table 2. The C1q-binding de novo donor-specific HLA antibodies (dnDSA), C3d-binding dnDSA, and IgG ≥5000 MFI dnDSA are associated with C4d+ antibody-mediated rejection (AMR). Table 3. Kidney transplant recipients with de novo donor-specific HLA antibodies (dnDSA) of IgG MFI ≥5000 but negative for C3d and C1q-binding. The median fluorescence intensity (MFI) of each dnDSA is given in parenthesis.
Table 3. Kidney transplant recipients with de novo donor-specific HLA antibodies (dnDSA) of IgG MFI ≥5000 but negative for C3d and C1q-binding. The median fluorescence intensity (MFI) of each dnDSA is given in parenthesis. In Press
15 Mar 2024 : Review article
Approaches and Challenges in the Current Management of Cytomegalovirus in Transplant Recipients: Highlighti...Ann Transplant In Press; DOI: 10.12659/AOT.941185
18 Mar 2024 : Original article
Does Antibiotic Use Increase the Risk of Post-Transplantation Diabetes Mellitus? A Retrospective Study of R...Ann Transplant In Press; DOI: 10.12659/AOT.943282
20 Mar 2024 : Original article
Transplant Nephrectomy: A Comparative Study of Timing and Techniques in a Single InstitutionAnn Transplant In Press; DOI: 10.12659/AOT.942252
28 Mar 2024 : Original article
Association Between FEV₁ Decline Rate and Mortality in Long-Term Follow-Up of a 21-Patient Pilot Clinical T...Ann Transplant In Press; DOI: 10.12659/AOT.942823
Most Viewed Current Articles
05 Apr 2022 : Original article
Impact of Statins on Hepatocellular Carcinoma Recurrence After Living-Donor Liver TransplantationDOI :10.12659/AOT.935604
Ann Transplant 2022; 27:e935604
12 Jan 2022 : Original article
Risk Factors for Developing BK Virus-Associated Nephropathy: A Single-Center Retrospective Cohort Study of ...DOI :10.12659/AOT.934738
Ann Transplant 2022; 27:e934738
22 Nov 2022 : Original article
Long-Term Effects of Everolimus-Facilitated Tacrolimus Reduction in Living-Donor Liver Transplant Recipient...DOI :10.12659/AOT.937988
Ann Transplant 2022; 27:e937988
15 Mar 2022 : Case report
Combined Liver, Pancreas-Duodenum, and Kidney Transplantation for Patients with Hepatitis B Cirrhosis, Urem...DOI :10.12659/AOT.935860
Ann Transplant 2022; 27:e935860








